Thank you for visiting nature.com. You are using a browser version with limited support for CSS. To obtain the best experience, we recommend you use a more up to date browser (or turn off compatibility mode in Internet Explorer). In the meantime, to ensure continued support, we are displaying the site without styles and JavaScript.
- View all journals

Bioremediation articles from across Nature Portfolio
Bioremediation is a process that uses living organisms, mostly microorganisms and plants, to degrade and reduce or detoxify waste products and pollutants.
Latest Research and Reviews
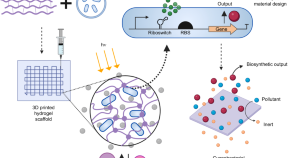
Phenotypically complex living materials containing engineered cyanobacteria
Engineered living materials (ELMs) are emerging as a field at the intersection of materials science and synthetic biology. Here, the authors describe a photosynthetic ELM composed of genetically engineered cyanobacteria in a hydrogel matrix, capable of bioremediation and inducible cell death.
- Debika Datta
- Elliot L. Weiss
- Jonathan K. Pokorski
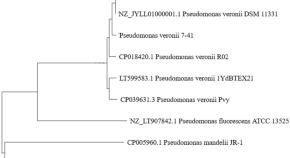

Pseudomonas veronii strain 7–41 degrading medium-chain n -alkanes and polycyclic aromatic hydrocarbons
- S. A. Mullaeva
- Ya. A. Delegan
- A. A. Vetrova
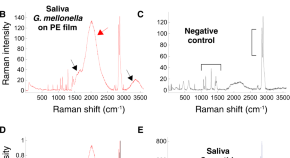
Wax worm saliva and the enzymes therein are the key to polyethylene degradation by Galleria mellonella
The crucial first step in the biodegradation of polyethylene plastic is oxidation of the polymer. This has traditionally required abiotic pre-treatment, but now Bertocchini and colleagues report two wax worm enzymes capable of catalyzing this oxidation and subsequent degradation at room temperature.
- A. Sanluis-Verdes
- P. Colomer-Vidal
- F. Bertocchini
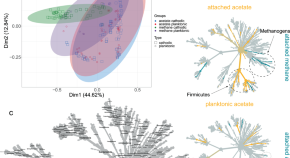
A meta-analysis of acetogenic and methanogenic microbiomes in microbial electrosynthesis
- Simon Mills
- Paolo Dessì
- Umer Zeeshan Ijaz
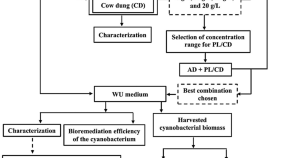
A waste-to-wealth initiative exploiting the potential of Anabaena variabilis for designing an integrated biorefinery
- Dipanwita Deb
- Nirupama Mallick
- P. B. S. Bhadoria

Effective treatment of aquaculture wastewater with mussel/microalgae/bacteria complex ecosystem: a pilot study
- Yongchao Li
- Weifeng Guo
News and Comment
A solution for mercury pollution.
Wu et al. provide mechanistic insight into how the plant symbiotic fungus Metarhizium robertsii can reduce mercury accumulation in soil.
- Michael Attwaters
Supporting the strategic pillars of translational research in biofilms
- Miguel Camara
- Alain Filloux
Bottlenecks and opportunities for synthetic biology biosafety standards
The lack of innovative standards for biosafety in synthetic biology is an unresolved policy gap that limits many potential applications in synthetic biology. We argue that a massive support for standardization in biosafety is required for synthetic biology to flourish.
- Michele Garfinkel
- Markus Schmidt
Pesticides and soil viruses
A study investigating the microbiome of contaminated soil finds that auxilliary metabolic genes of viruses contribute to pesticide degradation.
- Ursula Hofer
Vying over spilt oil
This month's Genome Watch explores the diverse microbial community involved in the bioremediation of oil spills.
- Tu Anh Pham N.
- Blessing O. Anonye

Crop pests: Under attack
The threat of insects to agriculture is set to increase as the planet warms. What action can we take to safeguard our crops?
Quick links
- Explore articles by subject
- Guide to authors
- Editorial policies
- Reference Manager
- Simple TEXT file
People also looked at
Review article, microbial biodiversity and bioremediation assessment through omics approaches.

- 1 Department of Botany, Mohanlal Sukhadia University, Udaipur, India
- 2 Laboratory of Phytopathology and Microbial Biotechnology, Department of Botany, Mohanlal Sukhadia University, Udaipur, India
Industrialization, population burst, and changing lifestyles have resulted in the genesis of non-degradable pollutants languishing the environment and human health. Biological approaches using microorganisms are gaining importance as an eco-friendly and cost-effective substitute to mitigate the pollution load. Microorganisms can survive in a divergent environment and produce metabolites that can degrade and transform pollutants making it possible to revive contaminated sites naturally. Modern omics technologies like metagenomics, transcriptomics, proteomics, etc. have been used nowadays design strategies to study ecology and diversity of microorganisms and their application in environmental monitoring and bioremediation. The present article will focus on the omics techniques reportedly used in environmental monitoring to tackle the pollution load.
Introduction
Pollution load is increasing at an alarming rate as a result of industrialization and population outburst. The industrialization has caused in the utilization and production of chemicals for hi-tech innovations which ensued the generation of non-biodegradable pollutants like xenobiotics, hydrocarbons, heavy metals, etc. ( Labie, 2007 ). These toxic pollutants remain persistent in the environment and pose a serious threat to living organisms. Increasing awareness has generated numerous approaches using advanced scientific technology to audit and curtail this arduous global issue. The widely accepted technique for decontaminating a polluted environment in an eco-friendly and sustainable manner is bioremediation ( Paul et al., 2005 ; Raghunandan et al., 2018 ).
Bioremediation is an enticing process to detoxify environmental contaminants. Microorganisms, such as fungi, yeast, and bacteria have been considered to be outstanding organisms for detoxification of pollutants ( Zhong and Zhou, 2002 ; Luciene et al., 2015 ; Abou Seeda et al., 2017 ). It assures a cheap, simple, and eco-friendly cleanup method ( Lovley, 2003 ). Microorganisms are nutritionally flexible and have the flair to adapt to extreme environmental conditions. They also possess numerous intracellular and extracellular enzymes which utilize the complex pollutants and convert them into carbon and energy source ( Nojiri and Tsuda, 2005 ; Thakur et al., 2019 ). They also undergo a rapid genetic transformation which enables them to acquire new metabolic routes for deterioration of xenobiotics ( Poirier et al., 2013 ; Igiri et al., 2018 ).
Microbial bioremediation strategies can be either ex-situ or in-situ . Ex-situ techniques consist of transporting pollutants from polluted sites to another site for treatment while in situ techniques treat polluted substances at the site itself. Ex-situ remediation technique owing to its limitations is not considered a choice of cleanup by many researchers. It may or may not be lucrative at particular sites and may be possible that the microorganisms which assisted in clean-up of pollutants under in-vitro conditions fail to remove them effectively under in-vivo conditions ( Head et al., 2003 ; Barupal et al., 2019a , b ). The mode of action and growth of microorganisms in polluted sites needs to be more studied for a better understanding ( Lovley, 2003 ). Thus, extensive research needs to be focused on elementary and applied facets of microbial synergism to assess the biodegrading capacity microorganisms present in polluted sites and the changes in their structure and metabolic pathways.
Microbial bioremediation strategies are based on the consortia of diverse organisms that are indigenous to the polluted sites and possess immense metabolic potential. Isolating and purifying such indigenous microorganisms gives some knowledge of the microbial metabolites and the degradation processes. The method to access the microbial world remains immensely perplexing, as the majority of microbes in the environment are non-culturable under in-vitro conditions ( Handelsman, 2004 ). Only a petite percent of microorganisms from diverse environment samples are actively culturable effortlessly and thus not accessible for fundamental research ( Dickson et al., 2014 ; Bursle and Robson, 2016 ; Awasthi et al., 2020 ).
The recent advances in molecular tools enabled a cutting edge to assess the unculturable organisms from natural environments ( Handelsman, 2004 ; Gilbert and Dupont, 2011 ). Molecular approaches like genomics, proteomics, transcriptomics, metabolomics, fluxomics, etc. gave more vision about the microbial communities inhabiting a particular environmental niche ( De Sousa et al., 2018 ; Gupta et al., 2020 ; Rodríguez et al., 2020 ). The onsets of next-generation sequencing methods and in-silico analyses have facilitated researchers to address the problems associated with unculturable microorganisms ( Maphosa et al., 2010a ; Malla et al., 2018 ). These technologies have abetted the collection exhaustive biological data about microorganisms, their cellular mechanisms, and biomolecules like enzymes associated with bioremediation ( Zhang et al., 2010 ; Misra et al., 2018 ; Pandey et al., 2019 ).
Bioremediation holds future assurance for detoxifying the contaminated sites. However, the technique is still riddled with complications that need to be addressed. The most critical step is to comprehend the indigenous microbiome, their synergy with the environment, and its associated factors which help in the expression of their genotype ( Chakraborty et al., 2012 ; Callaghan, 2013 ). Hence, a multidisciplinary approach is required to unveil the pathways and chemistry that remains hidden for making bioremediation a versatile technique for environmental monitoring. Earlier published research papers have emphasized on individual genomic approaches in-depth for microbe mediated bioremediation. Hence, the present review aims to discuss a brief overview of the prevalent multi-omics techniques to investigate and understand the structural and functional aspects of the microbial community involved in bioremediation. This review article also provides a comparative analysis of commonly employed sequencing platforms and reviews a few recent signs of progress of integrated omics approaches to analyze complex microbial communities in diverse environmental samples. Therefore, the present article will discuss the modern and effective method of “omics” technologies for environmental monitoring and bioremediation.
Genomics and 16s rRNA for Bioremediation
The emergence of genomic technologies has boosted the treatment of contaminated environments in a sustainable manner. Cultivation-independent analyses of microorganisms from contaminated sites have amplified the perception of community dynamics and myriad of microbes actively involved in bioremediation ( Kapley et al., 2007 ; Desai et al., 2010 ). The identification of the microbial communities using modern genomic tools has enabled the detection of distinctive microorganisms that were not approachable by culture-based techniques. Gene amplification (using PCR) and sequencing techniques have proven exceptionally useful in evaluating the microbial community ( Malik et al., 2008 ; Rani et al., 2008 ; Gołebiewski and Tretyn, 2020 ).
The discovery of a highly conserved and variable gene sequence, 16S rRNA in all microorganisms is considered as the highest quality level for describing phylogenetic similarities among organisms in microbial communities ( Lovley et al., 1991 ; Lovley, 2003 ). 16S rRNA gene sequence analysis can be used for a complete assessment of microbial diversity by selectively amplifying and sequencing the hypervariable regions of the 16S rRNA gene. It is a highly efficient and cost-effective technology easily accessible by various bioinformatics tools and has become a frequently used technique for profiling intricate microbial communities ( Han et al., 2020 ). It can be used to identify novel, unculturable, and phenotypically unidentifiable microbes ( Clarridge, 2004 ).
The phylogenetic organization of the microbes linked with bioremediation processes can be determined by investigating the 16S rRNA sequences obtained from contaminated environments ( Lovley, 2003 ; Rogers and McClure, 2003 ). 16S rRNA was used to elucidate the composition of microbial communities and the multifariousness of the dioxygenase genes in the soil of a coal tar mixing plant to study the genetics of PAH degradation ( Kumar and Khanna, 2010 ; Viant and Sommer, 2013 ; Sakshi Haritash, 2020 ). Kou et al. (2018) reported 16S rRNA gene amplicon sequencing to study the abundance and diversity of the microbial community in soil polluted with heavy metals like lead, zinc, and copper in Shanghai. 16S rRNA gene sequencing along with membrane fatty acid profile was used to identify soil bacterium Pseudomonas species capable of degrading polyurethane from a site containing an abundance of fragile plastic waste ( Cárdenas Espinosa et al., 2020 ).
Microarrays
DNA microarray is widely known as a DNA chip or a biochip is one of the most promising methods in functional genomics. It is an assortment of microscopic DNA spots deposited or synthesized in a two dimensional or three-dimensional arrays on a solid surface like glass, silicon chips, or nylon membrane by covalent or non-covalent interactions. It allows the analysis of multiple genes at once without PCR amplification of the individual genes. DNA microarrays have been reported to be used to assess the physiology and catabolic gene expression profile of microorganisms isolated from environmental samples ( Schut et al., 2001 ; Dennis et al., 2003 ; Chandran et al., 2020 ). Rhee et al. (2004) established an extensive 50-mer-based oligonucleotide microarray to identify genes associated with biodegradation and biotransformation in microbial communities. DNA microarrays were used in transcriptional profiling, quantify genes in biological samples, stress gene analysis of microbial genomes, etc. ( Cho and Tiedje, 2002 ; Muffler et al., 2002 ; Greene and Voordouw, 2003 ; Chittora et al., 2020 ). Various microarrays designed to examine microbial communities are discussed below.
PhyloChip Arrays
It is an ease quality chip microarray created by the business head Affymetrix to recognize different bacterial and archaeal organisms from complex microbial communities. It gives a progressively quick, complete, and precise testing method for samples retrieved from the environment without the use of any culture techniques. These chips contain large information of genes and are widely used in hybridization-based identification and study of mutations and polymorphisms like single nucleotide polymorphisms or disease- pertinent mutations analysis. They have been used to study microbial profile in extreme biological systems like sun powered salterns, industrial effluents, olive-mill squanders, coral reefs, etc. ( Nikolaki and Tsiamis, 2013 ). Phylochip arrays have been reportedly used to explore the bacterial diversity and community structure in chromium contaminated industrial wastewater ( Katsaveli et al., 2012 ), Australian sugar cane processing plants ( Sharmin et al., 2013 ).
Functional Gene Arrays (FGAs)
They are the particular kind of DNA microarrays incorporating probes for pivotal genes associated with microbial functional progressions ( Cho and Tiedje, 2001 ; Palmer et al., 2006 ). FGAs are useful for evaluating the functional composition and network of microbial communities ( Zhou et al., 2015 ; Shi et al., 2019 ). It is commonly employed for the synchronized inspection of multiple functional genes ( Greene and Voordouw, 2003 ; Palmer et al., 2006 ). These arrays are widely used to study microbial communities from environmental samples because the majority of microbes in such samples are highly diverse, metabolically flexible, and unculturable ( van Nostrand et al., 2011 ).
GeoChip Arrays
It is the most extensive DNA microarray accessible for contemplating microbial communities related to biogeochemical cycling, ecosystem management, environmental restoration, and remediation, etc. It can be utilized to study microbial association in soil, water, air, human and animal sources. It does not entail any preceding information of the microbial community and can detect less abundant microorganisms preventing annotation bias. The process is comparatively fast, and uses either DNA or RNA and utilizes 56,990 gene sequences from 292 functional genes consuming 27,812 probes. GeoChip uses key enzymes or genes to spot various microbe mediated mechanisms for biogeochemical cycles of nitrogen, sulfur, phosphorus, etc. resistance mechanisms for heavy metals like silver, cadmium, copper, mercury, nickel, lead, zinc, etc. ( van Nostrand et al., 2011 ; Zehra et al., 2020 ), degradation pathways to identify and track the degradation of xenobiotics frequently found in the environment ( He et al., 2010 ; Xie et al., 2011 ). It has been reported as an efficient, sensitive, perceptive and quantitative tool for investigating the functional structure of microbial communities ( Zhou et al., 2008 ; Xue et al., 2016 ) from divergent environments like soils, aquatic systems, extreme environments, contaminated habitats, etc. ( He et al., 2018 ; Shi et al., 2019 ).
Omics Approaches
Advances in molecular techniques like proteomics, transcriptomics, metagenomics, etc. have opened new strategies and opportunities in environment management ( Plewniak et al., 2018 ; Rodríguez et al., 2020 ). These methods have accelerated the study of microbial community structure which was earlier dependent on culture technologies ( Iwamoto and Nasu, 2001 ; Gutleben et al., 2018 ). It possesses the potential to evaluate the genetic diversity of environmentally pertinent microorganisms and analyze novel functional genes related to the catabolism of pollutants ( Eyers et al., 2004 ; Meena et al., 2019 ). Figure 1 depicts the various types of omics approaches commonly used for microbial community analysis.
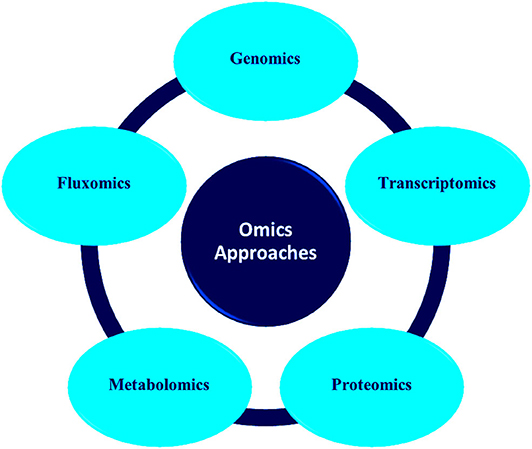
Figure 1 . Various types of omics approaches for microbial communities analyses.
Omics technology is the molecular biological approach, which facilitates the analysis of biomolecules like DNA, RNA, proteins, and metabolites from individual organisms and the whole community at the same time ( Gutierrez et al., 2018 ). To study gene regulation in the anthropogenic environment, mRNA expression (transcriptomics), and whole community expression (metatranscriptomics) can be studied ( Schneegurt and Kulpa, 1998 ; Seo et al., 2009 ; Roume et al., 2015 ). Anthropogenically induced proteins from the contaminated sites can be tracked by studying the proteins using metaproteomics ( Desai et al., 2010 ). Metagenomics analyzes the microbial communities directly from the ecological environment which in turn reduces the labor of screening and culture of microbes ( Riesenfeld et al., 2004 ; Panagiotou et al., 2007 ; Ghazanfar et al., 2010 ). The study of protein through which anthropogenically induced proteins can be tracked from the contaminated sites is called metaproteomics ( Desai et al., 2010 ). The analysis of primary and secondary proteinaceous metabolites synthesized by the organisms during distinct environmental situations is metabolomics ( Desai et al., 2010 ). Multi-omics approaches are commonly reported to study microbial communities because a single omics analysis cannot disclose the functional activity of the microbial community ( Shah et al., 2013 ; Meena et al., 2018 ).
Metagenomics
A major share of the microorganisms in the environment is non-culturable under laboratory conditions ( Rashid and Stingl, 2015 ). Metagenomics can be used to explore such non-culturable microbes thriving in different environments ( Oulas et al., 2015 ; Bilal et al., 2018 ). Metagenomic sequences reveal DNA sequences of uncultured microbes thriving in the environment which can be used for new biotechnology applications. The metagenomic information will enable researchers to integrate pure culture study with genomics ( Hodkinson and Grice, 2015 ). It uses the pool of environmental genomes of microorganisms which increases the probability to discover unique genes and diverse pathways with new enzymes containing highly specific catalytic properties ( Scholz et al., 2012 ; Yergeau et al., 2017 ; Awasthi et al., 2020 ). Due to the dearth of high-throughput sequencing technologies and software early metagenomic studies were focused on environments like acid mine drainage and the human gut microbiome. With the advancement of powerful software tools and molecular technologies extreme environments with extreme temperature, alkalinity, acidity, low oxygen, deep-sea hydrothermal vents, heavy metal polluted soils, etc. have been studied using metagenomics which provides infinite chances for bioprospecting and exploring of novel biomolecules like proteins, enzymes, etc. ( Oulas et al., 2015 ; Awasthi et al., 2020 ).
Metagenomic assessment analysis commences with the isolation of nucleic acid (DNA or RNA) from the environmental samples. Dynamic microbial populaces in polluted environments can be evaluated by genome improvement monitored by metagenome investigation ( Chen and Murrell, 2010 ). Stable Isotope Probing (SIP) cataloging can be utilized to specifically enhance the RNA, DNA, or phospholipids of dynamic microbial populaces. Metagenomic reference libraries are prepared by cloning DNA fragments isolated from the environmental samples in an appropriate vector [phage, plasmid, bacterial artificial chromosome (BAC)] and then reconstructed into a host bacterium. The consequential transformants can be screened for phylogenetic markers, conserved genes, expression of specific traits, etc. ( Handelsman, 2004 ). Biological information from metagenomic libraries can be acquired based on two types of analysis: function-driven or sequence-driven analysis. The function-driven analysis is based on the identification of clones that express their functional activity while sequence driven analysis is based on the sequencing of clones with phylogenetic anchors or conserved DNA sequences that is the plausible origin of the DNA fragment ( Wu et al., 2010 ; Felczykowska et al., 2015 ; Wong, 2018 ).
Sequence-based screening is increasingly used owing to the availability of several software for data analysis and the ease to assess metagenomic sequencing data. This approach is predominantly influenced by the precision of genome annotation, the integrity of the available data, algorithms, and facts in databases to ascertain the function of novel genes ( Ferrer et al., 2009 ). It may not be useful if the sequence analogy does not complement to a functional association or the original gene has less analogy to some genes whose products have been investigated biochemically or a specific gene is capable to accomplish diverse tasks in the cell ( Hallin et al., 2008 ). In such cases, function-driven screening is preferred to discover genes with novel functions or to explore the sequence variation of protein families ( Singh et al., 2009 ; Meena et al., 2016 ). The flowchart in Figure 2 represents the general procedures adopted for metagenomic research.
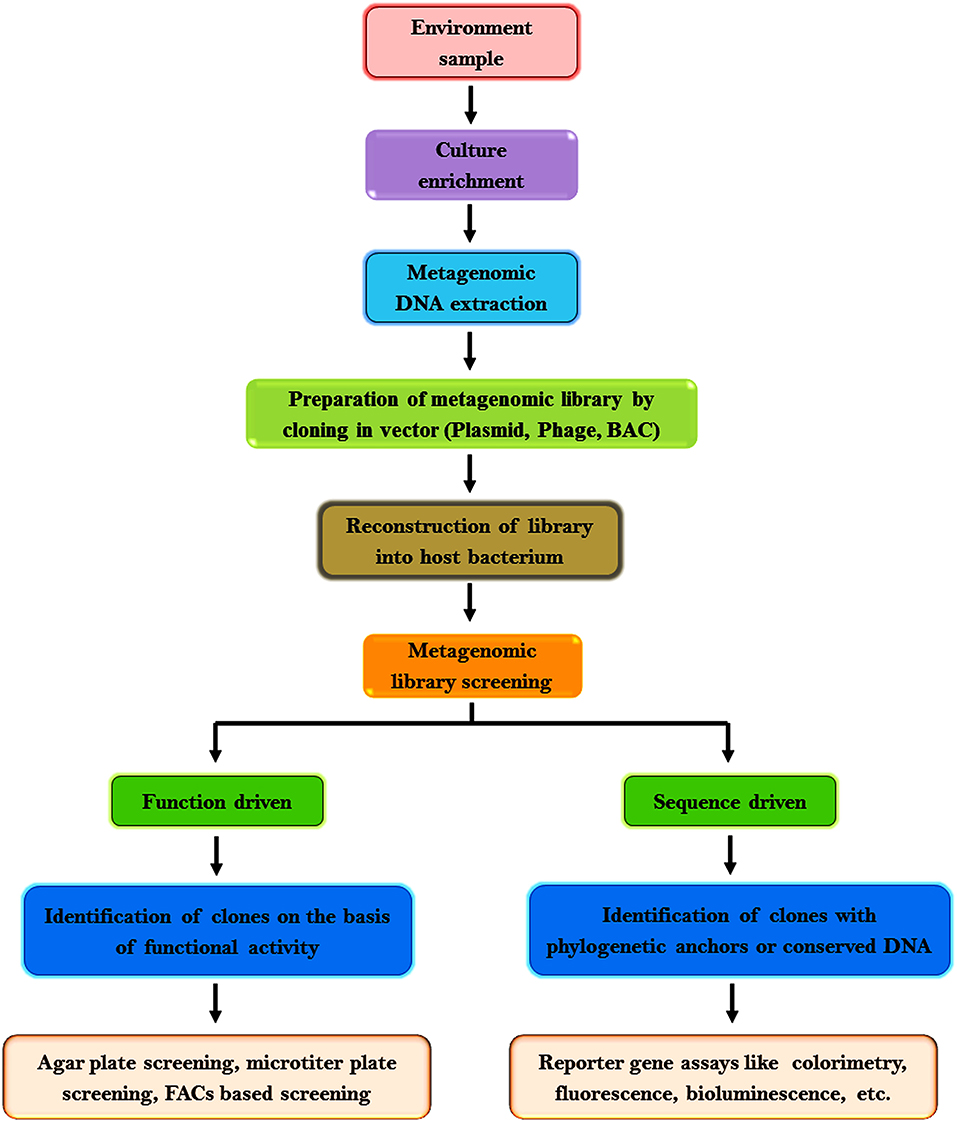
Figure 2 . Workflow of metagenomics research.
Metagenomic analysis has opened new frontiers to analyze microbial communities, their genetic diversity, and metabolic pathways. It has provided opportunities to discover microbial consortia and genes involved in the bioremediation of xenobiotic compounds. Phenol degrading pathways of uncultivated bacteria in activated sludge was studied using metagenomics ( Sueoka et al., 2009 ). The metagenomic approach was used to characterize genes and metabolic pathways associated with the degradation of phenol and other aromatic compounds in sludge from a petroleum refinery wastewater treatment system ( Silva et al., 2013 ). Vigneron et al. (2017) accomplished a comparative metagenomic study to understand the community structure and diverse functions of marine seep ecosystems in the Gulf of Mexico and reported that microorganisms in deep cold seeps have the potential to consume a range of non-methane hydrocarbons. Metagenome sequencing was employed to outline the functional potential, taxonomic community composition, and to predict the breakdown of chemical compounds of soils with organophosphorus pesticide exposure ( Jeffries et al., 2018 ). Metagenomic data analysis of microbial consortium emerging on diesel revealed the bacterial genera, genes, and enzymes involved in diesel degradation ( Garrido-Sanz et al., 2019 ).
Gaytán et al. (2020) combined physical and chemical analysis with metagenomics to explicate probable metabolic pathways associated with polyurethane-degrading to alleviate plastics and xenobiotics pollution. The persistent impact of petroleum pollutants on the taxonomic and metabolic structure of microbial mats were studied using metagenome and enriched mRNA metatranscriptome sequencing ( Aubé et al., 2020 ).
Metagenome Screening Methods
It can be screened using an activity-based (function-driven) or genome-driven (sequence-driven) approach ( Desai et al., 2010 ). A small insert library can be used to discover a single gene or small operons enciphering new metabolic function while large insert libraries are more preferred for a large number of gene groups or DNA fragments ( Daniel, 2005 ; Meena et al., 2015 ). Commonly employed activity-based screening strategies to acquire metagenome derived biocatalysts are agar plate screening, microfluidics-based screening, FACs based screening, microtiter plate screening, etc. It is based on the generation of a zone of clearance/inhibition, production of secondary metabolites (pigments, antibiotics, etc.), chromophore or fluorophore by the clones ( Popovic et al., 2017 ; Ngara and Zhang, 2018 ).
In genome driven approach reporter gene-based in vivo screening and selection strategies are employed. The ease in accessing metagenomics sequencing data and the availability of multiple softwares makes this approach in demand for exploring metagenomic libraries ( Lin et al., 2006 ; Dai et al., 2018 ). Genome driven approach is dependent on reporter gene present in the clones which may be colorimetric (LacZ), fluorescent (GFP), bioluminescent (LuxCDABE), conditional survival (CAT), acidification (AraBAD), ice nucleation (InaZ), motility (CheZ), cell display (LamB), etc. The commonly employed reporter based strategies are transcriptional regulator based (synthetic/natural), ribozyme based, post-translational modified reporter-based and fusion-based strategy. Among the reporter based strategies transcriptional regulator based approach is the most employed selection strategy ( Dietrich et al., 2010 ; van Rossum et al., 2013 ).
The effectiveness of sequence-based approaches relies on precise genome annotation and available data ( Ferrer et al., 2009 ). It uses algorithms and databases to conclude the functions of novel genes. Thus, these approaches may lag if the sequence analogy does not complement to a functional relationship, or if the novel gene has a feeble resemblance to any genes whose products have been analyzed biochemically, or a specific gene is competent to perform multiple functions in the cell ( Ferrer et al., 2009 ). Function-driven screening is the chosen method for discovering novel genes or for probing the sequence diversity of protein families ( Singh et al., 2009 ; Zhuang et al., 2010 ; Bier et al., 2018 ).
Genome Sequencing Technologies
Complete genome sequencing progressed through three technical (mechanical) transformations: first-generation sequencing (complete genome shotgun sequencing), next-generation sequencing (high throughput sequencing), and the third-generation sequencing (single molecule long-read sequencing) ( Loman and Pallen, 2015 ).
First generation sequencing
Frederick Sanger and Allen Maxam—Walter Gilbert sequencing techniques were categorized as the first generation DNA sequencing methods ( Liu et al., 2012 ). Sanger sequencing uses denatured DNA template, radioactively labeled primer, DNA polymerase, and chemically modified nucleotides called di-deoxynucleotides to generate DNA fragments with various lengths. Size of the DNA fragment is determined by the incorporated dNTPs. The DNA fragments are detached based on their lengths on gel electrophoresis and can be viewed by an X-ray or UV-light imaging system ( El-Metwally et al., 2014 ; Meena et al., 2017a , b ). This technique is expensive, time-consuming and do not permit the sequencing of complex genomes hence, it is widely utilized for single or low-throughput DNA sequencing nowadays ( Sanger and Coulson, 1975 ; Kchouk et al., 2017 ). Maxam-Gilbert sequencing is recognized as the chemical degradation method because it uses chemicals to cleave nucleotides. Chemical treatment generates breaks in nucleotide bases which lead to the generation of a series of marked fragments that can be detached based on their length/size by gel electrophoresis. It is considered hazardous owing to the usage of noxious and radioactive chemicals ( Maxam and Gilbert, 1977 ). These techniques were the most routinely used sequencing techniques by researchers till the rise of new sequencing technologies which opened new frontiers for genome analysis.
Next generation sequencing
The second-generation sequencing technology speeded up the process of sequencing by generating sequences with very high throughput at a low cost. It produces a massively parallel analysis from multiple samples where small sections of DNA are ligated with adaptors for random reads during DNA amplification which provides enormous data within a short duration ( Zhang et al., 2011 ). Next-generation sequencing technology involves library preparation, sequencing, base calling, alignment to the established genome, and assorted annotation. Library preparation begins with the fragmentation of DNA into multiple fragments by sonication, enzymatic digestion, or transposase followed by ligation with adaptor sequences. The prepared library is then amplified using clonal amplification and PCR methods to generate DNA replicas. DNA replicas are then sequenced using different approaches ( Samorodnitsky et al., 2015 ). The major platforms used for microbiome studies in next-generation sequencing are pyrosequencing (Roche/454 sequencing), Illumina, SOLiD, Ion Torrent, PacBio RS, etc. ( Hodkinson and Grice, 2015 ). Each sequencing platforms has its peculiar distinct advantages and disadvantages and selection of sequencing platform depend on the required read length, sequence accuracy, depth, and cost ( Table 1 ).
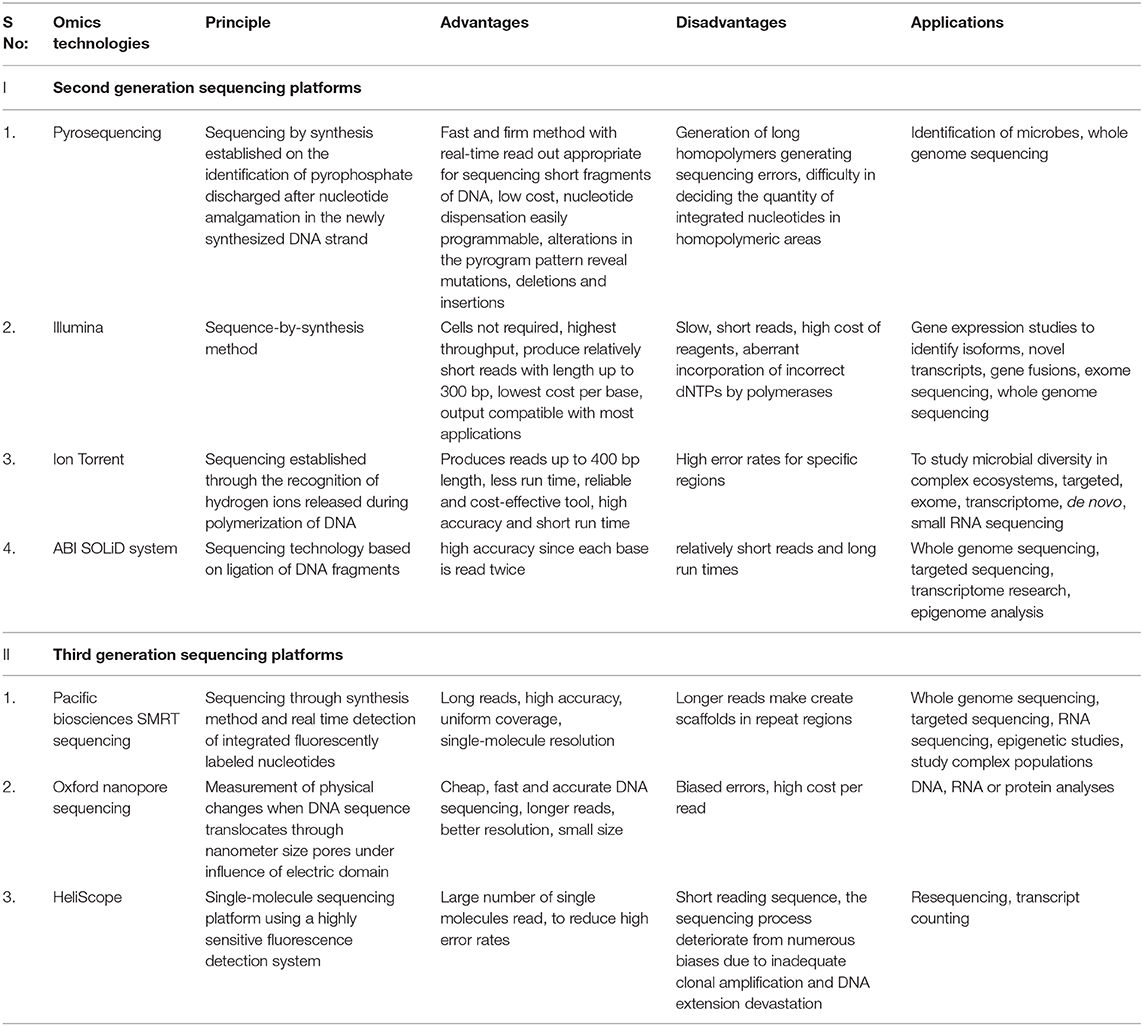
Table 1 . Comparative analysis of next generation sequencing for microbiome studies.
(i) Pyrosequencing technique
The pyrosequencing technique is sequencing by synthesis approach based on the detection of pyrophosphate release after the addition of a nucleotide to the newly synthesized DNA strand ( Mardis, 2008 ; Kchouk et al., 2017 ). Figure 3 shows the diagrammatic presentation of the pyrosequencing approach. DNA molecule is sheared and ligated with oligonucleotide adapters, affixed to a 28-μm bead, and amplified using PCR in an oil-water emulsion and pyrosequenced ( Ronaghi et al., 1996 ). The beads containing amplicon are then trapped in a picoliter sized well and pyrosequenced. The inclusion of non-labeled nucleotide results in enzyme-mediated liberation of inorganic pyrophosphate which can be detected computationally. Thus, repetitive pyrogenic cycles produce a DNA sequence with a read length of up to 1,000 base pairs and can produce ~1 million reads per run ( El-Metwally et al., 2014 ). The first bacterium to be sequenced through the pyrosequencing technique was a soil bacteria Myxococcus xanthus ( Vos and Velicer, 2006 ). Later, it has been employed to investigate microbial communities from divergent environment sources like freshwater, underground mine water, marine water, corals, fish, terrestrial animals, and mosquitoes ( Dinsdale et al., 2008 ).
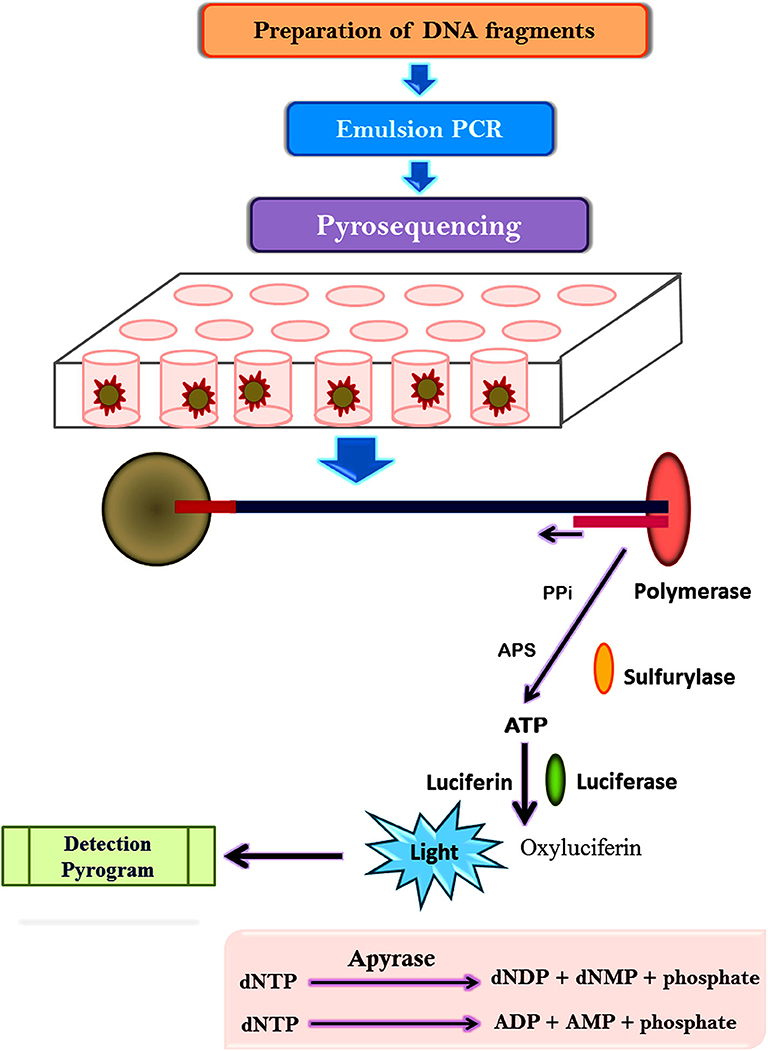
Figure 3 . Representation of pyrosequencing approach.
(ii) Sequencing by ligation on beads
It consists of multiple sequencing rounds. Figure 4 shows the ligase-mediated sequencing method of the Sequence by Oligonucleotide Ligation and Detection (SOLiD) sequencer (Life Technologies) and its step by step process. It begins with the attachment of adapters to DNA fragments fixed on 1 μm paramagnetic beads and amplified by PCR in an oil-water emulsion. PCR amplicons in beads were secured on a compact planar substrate and hybridized with a universal PCR primer paired with an adaptor. During the event of sequencing, the positional identity of the nucleotide is revealed by ligating universal primer to a fluorescently labeled DNA octamer. The cycle is continued till every base is sequenced two times which enhances the accuracy of this platform ( Shendure et al., 2005 ; Mardis, 2008 ; Meena et al., 2017c ) ( Figure 4 ).
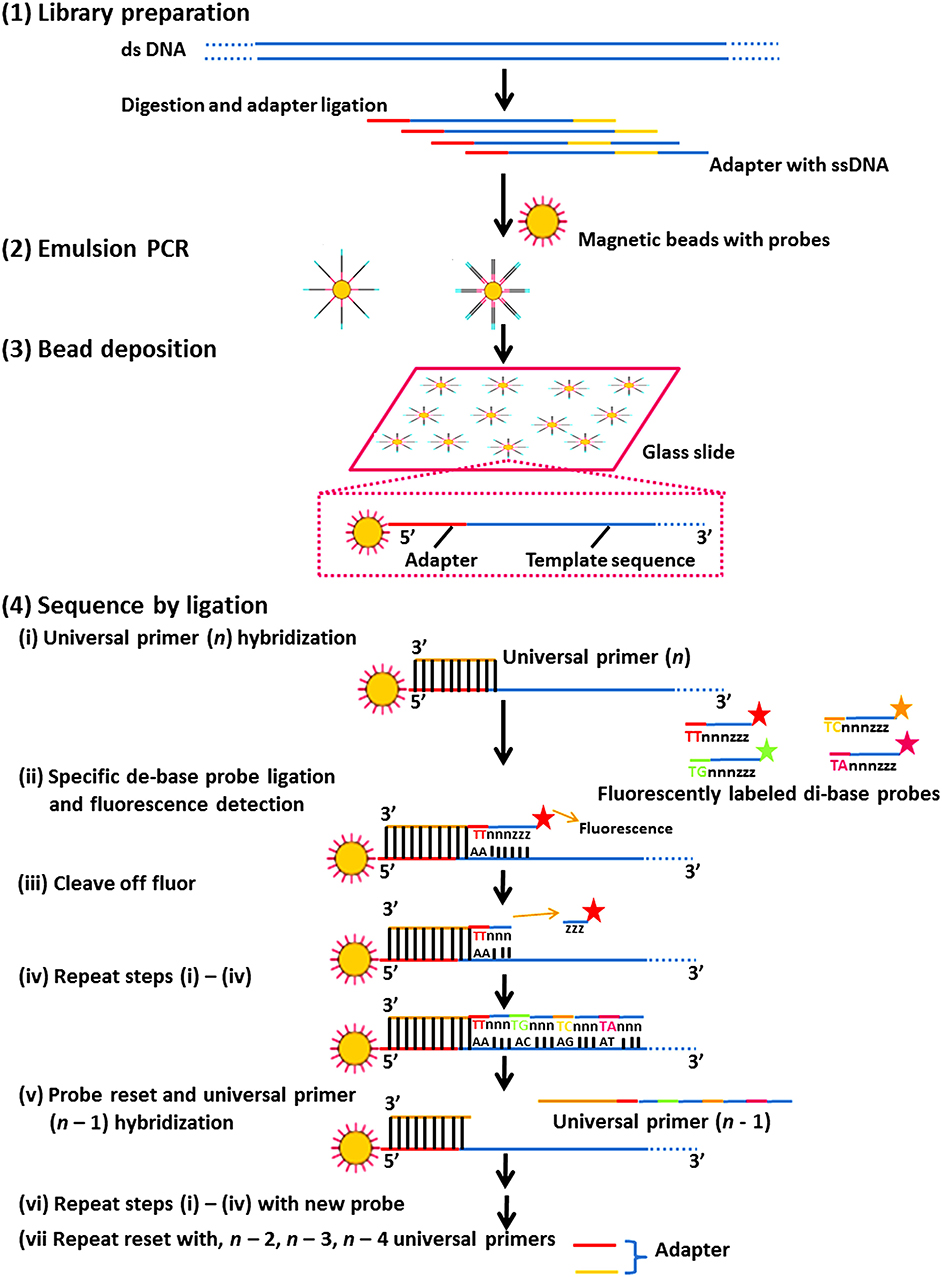
Figure 4 . The ligase-mediated sequencing approach of the Sequence by Oligonucleotide Ligation and Detection (SOLiD) sequencer (Life Technologies). (1) Library preparation: two distinctive adapters are ligated to sheared genomic DNA. (2) Emulsion PCR: emulsion PCR is managed with magnetic beads to generate “bead clones,” in which each comprises a solitary nucleic acid species. (3) Bead deposition: the beads are then associated to the outside surface of a glass slide. (4) Sequencing by ligation: ligase-mediated sequencing initiates by annealing a universal primer to the mutual adapter sequences on each amplified fragment (i), and subsequently DNA ligase is given accompanied by precise fluorescently labeled 8-mers, in which the two bases at the 3′ end of the probe are encoded by the appended fluorescent cluster. Every ligation step is monitored by fluorescence recognition (ii), after which a repossession step eliminates the bases from the ligated 8-mer (counting the fluorescent cluster) (iii), and correspondingly readies the extended probe for additional round of ligation (iv–vii). Subsequently every fluorescent cluster on a ligated 8-mer distinguishes a two-base amalgam, the consequential sequence peruses can be screened for base-calling errors vs. either correct polymorphisms or single base removals, by adjusting the individual states to a realized excellent reference sequence.
(iii) Illumina/Solexa sequencing
The DNA library preparation starts with random fragmentation of DNA into sequences followed by ligation of adapters to both ends of each sequence ( Adessi et al., 2000 ; Fedurco et al., 2006 ; Turcatti et al., 2008 ). Linkers are used to attach forward and reverse primers with complementarity to the adaptor on a glass surface. Bridge PCR intensifies the DNA fragment resulting in a “cluster” with multiple copies of a single DNA template. Each cluster comprises around one million replicas of the initial sequence. Nucleotide in the sequences is determined using sequencing by synthesis approach using reversible terminators where four amended nucleotides, sequencing primers, and DNA polymerases are incorporated so that the primers are hybridized to the sequences ( Meena and Samal, 2019 ).
It employs modified dNTPs containing a terminator with a fluorescent label at the 3′-OH end that blocks polymerization and allows only a single base addition to each growing DNA strand. Sequencing proceeds in cycles and single bases are added to all templates identically in each cycle, followed by the chemical cleavage of the fluorescent terminator which recognizes the incorporated nucleotide ( Bahamonde et al., 2016 ; Arazoe et al., 2018 ). Lasers are passed onto the DNA clusters to trigger the fluorescent label incorporated in the nucleotide base which are further detected and recorded by a camera. Multiple computer softwares are employed to interpret these signals into various nucleotide sequences. The process proceeds with the exclusion of the fluorescently-labeled terminator group with the incorporation of a new fluorescently-labeled terminator base until billions of clusters have been sequenced ( Bentley et al., 2008 ; Reuter et al., 2015 ). Illumina sequencing is extremely precise because the DNA sequence is explored base-by-base.
(iv) Ion torrent sequencing
It is analogous to pyrosequencing technology. Instead of using fluorescent-labeled nucleotides this approach relies on the release of a proton when a dNTP is added to DNA polymer. Adaptors are added to the fragmented DNA or RNA, and one molecule is placed onto beads and amplified by emulsion PCR. Each bead is laid into a single well of a slide containing single species of dNTPs, polymerase, and buffer. Nucleotide incorporation into DNA strand by polymerase releases hydrogen ion as a byproduct that decreases the pH. The decrease in pH is sensed by a pH sensor at the base of the microwell and transformed into a voltage which is proportional to the number of nucleotides incorporated. It is proficient to produce reads lengths of 200, 400, and 600 bp.
Third generation sequencing
Third generations sequencing offers lower sequencing charge and contented sample preparation without PCR amplification. The two most widely used sequencing platforms in third generations sequencing are Pacific Biosciences, Oxford Nanopore technology, and Heliscope technology. Pacific Biosciences developed the first genomic sequencer using single-molecule real-time technology (SMRT) approach ( Eid et al., 2009 ; Meena et al., 2017d ). This approach makes use of fluorescent labeling like other sequencing technologies. It detects nucleotide signals in real-time. It employs a framework made of several SMRT cells which comprises microfabricated nanostructures called zero-mode waveguides (ZMWs) ( Rhoads and Au, 2015 ). The bases of each ZMW contain DNA polymerase which binds the target DNA and starts replication. Fluorescently labeled nucleotides with different colors are then added to the reaction wells. With the enzymatic incorporation of each base, a flash of light is generated that recognizes the base and analyzed repetitively to produce the DNA sequence ( Rhoads and Au, 2015 ). It can produce a read length of about 1,500 bp.
Oxford nanopore technology uses electrophoresis to pass the DNA/RNA molecule through a nanopore. It makes use of electrolytic solutions and the utilization of a perpetual electric field. In this technology, double-stranded DNA is sheared and blunt-ended DNA molecules are formed with a termination repair stage. Two adaptors (Y adapter and a hairpin adaptor) conjugated with special motor protein are then added to the DNA which helps in unzipping the double-stranded DNA at the Y adapter and moves the DNA as a single strand through the nanopore. As the nucleic acid moves through the nanopore motor protein action generates a difference in ionic current due to the mobile nucleotides filling the pore. The deviation in ionic current is documented on a graphic model and then elucidated for sequence identification ( Jain et al., 2016 ; Lu et al., 2016 ).
HeliScope is another technology platform utilizing an extremely perceptive fluorescence detection system for single DNA molecule sequencing. DNA strands are fragmented using restriction enzymes detected by the addition of poly-A tail. The DNA fragments are hybridized to the flow cell plate with billions of oligo(dT) chains bound to its surface to produce an array of a primer annealed single DNA templates. Labeling is accomplished in “quads” comprising of 4 cycles each for each of the four nucleotide bases. Fluorescently labeled bases are added one at a time resulting in a template-dependent extension. A laser light illuminates the label, taking a read of the strands that have taken up a specifically labeled base which is further detected and recorded by a camera. Various computer programs interpret these signals into a nucleotide sequence. The label is then cleaved, and the next cycle proceeds with a new base ( Braslavsky et al., 2003 ; Meena et al., 2017e ).
Shotgun sequencing
Shotgun metagenomic sequencing is a powerful technique in microbial ecology because it provides a vigorous and reliable evaluation of microbial diversity ( Hillmann et al., 2018 ). It does not depend on PCR amplification and used to examine the functional potential and microbial composition of the community. It is the only way to study the microbial community with no markers like viruses ( Quince et al., 2017 ; Vermote et al., 2018 ). It allows strain-level remodeling in the taxonomic analysis and pathway predictions for the functional annotation of the microbiome under study ( Han et al., 2020 ).
It is an emerging molecular method to bridge the gap amid community structure and functional competence. It also helps in understanding the strategies adopted by microorganisms to thrive in adverse conditions ( Sharpton, 2014 ; Peabody et al., 2015 ; Ranjan et al., 2016 ). The functional activity of innate communities can be studied using the metagenomic shotgun sequencing approach. Figure 5 shows a schematic illustration signifying the standard metagenomic workflow. It has been used to new and existing microbial community, analyze new viruses ( Yozwiak et al., 2012 ; Quince et al., 2017 ; Vermote et al., 2018 ), uncultured bacteria ( Wrighton et al., 2012 ), unique proteins ( Godzik, 2011 ), and metabolic pathways ( Morgan et al., 2012 ).
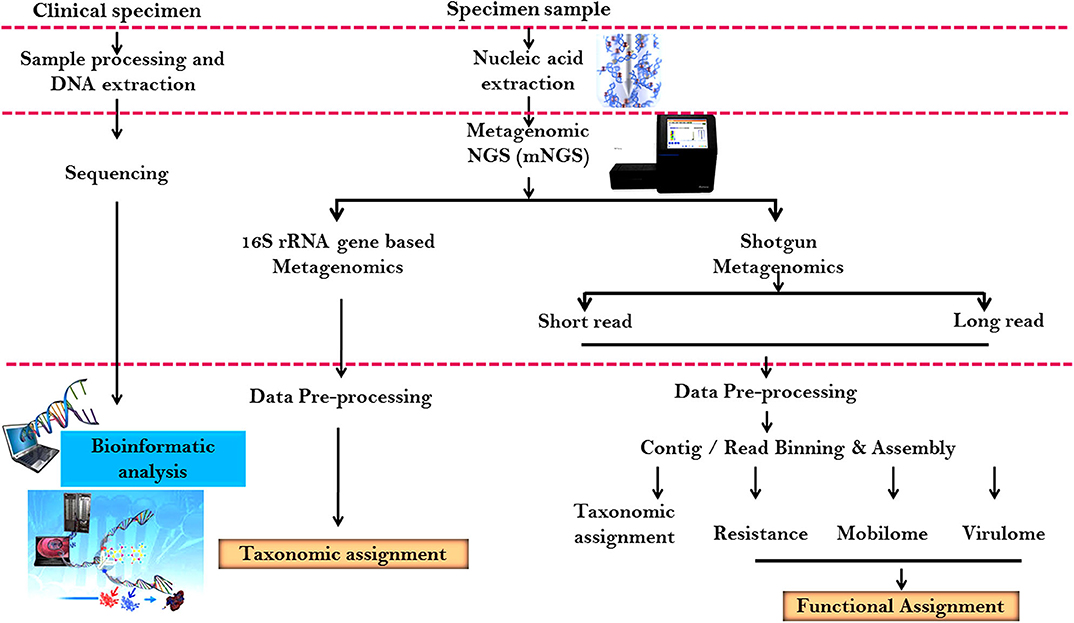
Figure 5 . Schematic illustration representing the standard metagenomic workflow.
Shotgun metagenomic sequencing of genomic DNA from a sample has equipped by library planning. The library planning workflow is analogous to consistent whole-genome sequencing, comprising of random DNA fragmentation and adapter ligation. Shotgun metagenomic sequencing workflow for taxonomy analysis consists of quality pruning and evaluation of a reference database involving whole genomes or specifically designed marker genes to create a taxonomy profile. Since, it contain all genetic information in a sample, the information can be used for supplementary analyses like metagenomic assembly and binning, metabolic function profiling, antibiotic resistance gene profiling, etc.
Whole-genome shotgun sequencing of environmental pooled DNA from seawater samples of Sargasso Sea near Bermuda elucidated gene content, diversity, and relative abundance (RA) of the organisms within these environmental samples ( Venter et al., 2004 ). Shotgun metagenomic sequencing of the cave microbiome in Thailand was studied to explore the taxonomic composition and metabolic potential of microorganisms in the soil ( Wiseschart et al., 2019 ). Shotgun metagenomic analysis of microbial communities from deep seabed petroleum seeps in the Eastern Gulf of Mexico revealed the presence of diverse communities of chemoheterotrophs and chemolithotrophs ( Dong et al., 2019 ). Whole-genome shotgun sequencing was engaged to identify the taxonomic diversity and gene repertoire of bacteria isolated from tannery effluents and petrol polluted soil samples for degradation of persistent organic pollutants like naphthalene, toluene, petrol, xylene, etc. ( Muccee and Ejaz, 2020 ).
Transcriptomics
The subdivision of genes transcribed in an organism is known as transcriptome. It is a potent network amid the proteome, genome, and cellular phenotype. It is also called gene expression profiling because it provides the understanding of up or down-regulation of genes under various environments in microbial communities. mRNAs analysis provides a direct vision of cell and tissue specific gene manifestation like (i) the existence, non-existence, and assessment of transcript, (ii) assessment of alternative splicing to foresee protein isoforms, and (iii) quantitative evaluation of genotype impact on gene expression via expression assessable trait loci analysis or allele-specific expression.
Microarray and sequencing techniques are employed for transcriptome analysis ( Maroli et al., 2018 ). Microarrays help to assess the expression of genes while RNA-sequencing uses next-generation sequencing to determine the amount of RNA in a sample. The RNA-microarray strategy uses pre-designed probes which make this approach cheaper, potent, and an enhanced technique for studying protein expression. RNA-sequencing is extensive as it facilitates different types of RNA at a much-advanced coverage and broad discovery studies ( Shendure, 2008 ; Nagalakshmi et al., 2010 ). The generation of raw transcriptome data involves purification of fine RNA of interest followed by transformation of RNA to complementary DNA (cDNA), fragmenting cDNA to build a library using sequence by synthesis (RNA-sequencing), running the microarray or sequence through superior software platform and carrying out ad hoc QC.
DNA microarray is a powerful technique in transcriptomics that supports in reviewing and evaluating mRNA expression of every single gene existing in an organism. The whole-genome DNA microarray technology has made it possible to define and investigate expressions of genes subjected to modulation with a fluctuating environment ( Gao et al., 2004 ). It has been employed to evaluate variance in metabolic and catabolic gene expressions, to analyze the microbial community physiology from diverse environments, identify new bacterial species, etc. ( Dennis et al., 2003 ; Greene and Voordouw, 2003 ).
Expression quantitative trait loci (eQTL) mapping is used as a persuasive strategy to explore and consolidate gene expression (RNA data) with genetic variants (DNA data). eQTL evaluation helps in identifying genetic variants that dominate mRNA expression levels. They furnish the relation between genetic variants and gene expression and can thus be used to exploit the fundamental molecular networks linked with a peculiar trait ( Yao et al., 2015 ). The regulation of gene expression can also be studied by the collective analysis of mRNA and microRNA levels. MicroRNAs (mRNAs) are short, non-coding RNA molecules that control transcription of mRNA. The precise binding of mRNAs to a target mRNA (by sequence homology) either impedes mRNA binding to the ribosome or targets it for degradation. Hence, it is therefore obvious that not only miRNA levels, but also their regulation by mRNAs are vital for an extensive gene expression dynamics ( Pritchard et al., 2012 ). mRNA profiling along with miRNA expression can be used to explore variations in the transcriptome profile, particularly to identify the miRNA transcripts that are subjected to regulation, emphasizing the probable molecular pathways supporting a particular trait or condition. This method has been, successfully used to explore gene expression in specific subtypes of gastrointestinal tumors ( Pantaleo et al., 2016 ), estimate modification of gene expression in wound-healing impaired fibroblasts from diabetic patients ( Liang et al., 2016 ).
Comparative transcriptomics revealed highly-upregulated degradation pathways and putative transporters for phenol to improve phenol tolerance and utilization by lipid-accumulating Rhodococcus opacus PD630 ( Yoneda et al., 2016 ). Transcriptome analysis of activated sludge microbiomes decoded the role of the nitrifying organisms in heavy oil degradation ( Sato et al., 2019 ). Transcriptome analyses of crude oil degrading Pseudomonas aeruginosa strains revealed the significance of differentially expressed genes implicated in crude oil degradation ( Das et al., 2020 ).
A proteome is the set of proteins formed inside a cell, tissue, organ, or organism. The branch of science that explores and study of proteomes is called proteomics. The proteomic analysis helps in decoding molecular mechanisms, metabolic pathways, post-translational modifications, etc. inside the cell. It has enabled the tracking and analysis of universal expression of proteins in microorganisms residing in contaminated areas due to human-induced activities ( Kim et al., 2004 ). Meta proteomics or community proteomics is the analysis of complete protein profile from microbial communities inhabiting in a specific environment ( Hart et al., 2018 ). It is also defined as a functional genomics approach because it helps in exploring the protein expression pattern of one organism and helps in generating a protein map of all expressed proteins by one organism thriving in a selected environment. Metaproteomic analyses are dependent on the effectiveness of proteins extraction, the techniques adopted for proteins fractionation from an intricate mixture, and the explicit identification of fractionated peptides/proteins, database search, and data interpretation ( Verberkmoes et al., 2009 ). Apart from this potent quantification methods are essential to relate expression patterns of proteins under different conditions. It relies on techniques like two-dimensional electrophoresis and mass spectroscopy to analyze the role of various microbial catabolic enzymes in biodegradation pathways ( Wilmes and Bond, 2006 ; Meena and Swapnil, 2019 ).
For analyzing the proteome of the microbial communities from random environments following steps have been developed. Figure 6 shows the highlights of the general workflow of proteomic analysis for microbial community studies. Environmental samples being complex requires unambiguous strategies for sample collection or protein extraction. Two strategies have been developed for protein separation and identification. One strategy is based on one-dimensional or two-dimensional polyacrylamide gel electrophoresis (2-D PAGE). Proteolytic enzymes like trypsin is used to digest the excised targeted protein bands to peptides which are later analyzed by mass spectrometry or tandem MS analysis, database probing, and bioinformatic analysis ( Wang et al., 2014 ). The second strategy is based on liquid chromatography coupled with MS/MS (LC-MS/MS) where proteases are used to digest the entire proteome into a complex peptide mixture and are purified by cation exchange chromatography or microcapillary reverse-phase. This method increases the proteome coverage, allows high-throughput recognition of proteins within a short duration ( Verberkmoes et al., 2009 ), and allows detection of insoluble membrane proteins ( Wu and Yates, 2003 ). The liquid chromatography centered methodology approach has become the cascade of microbial community proteomic studies. The disadvantage of this strategy is reproducibility, dynamic range, and database availability.
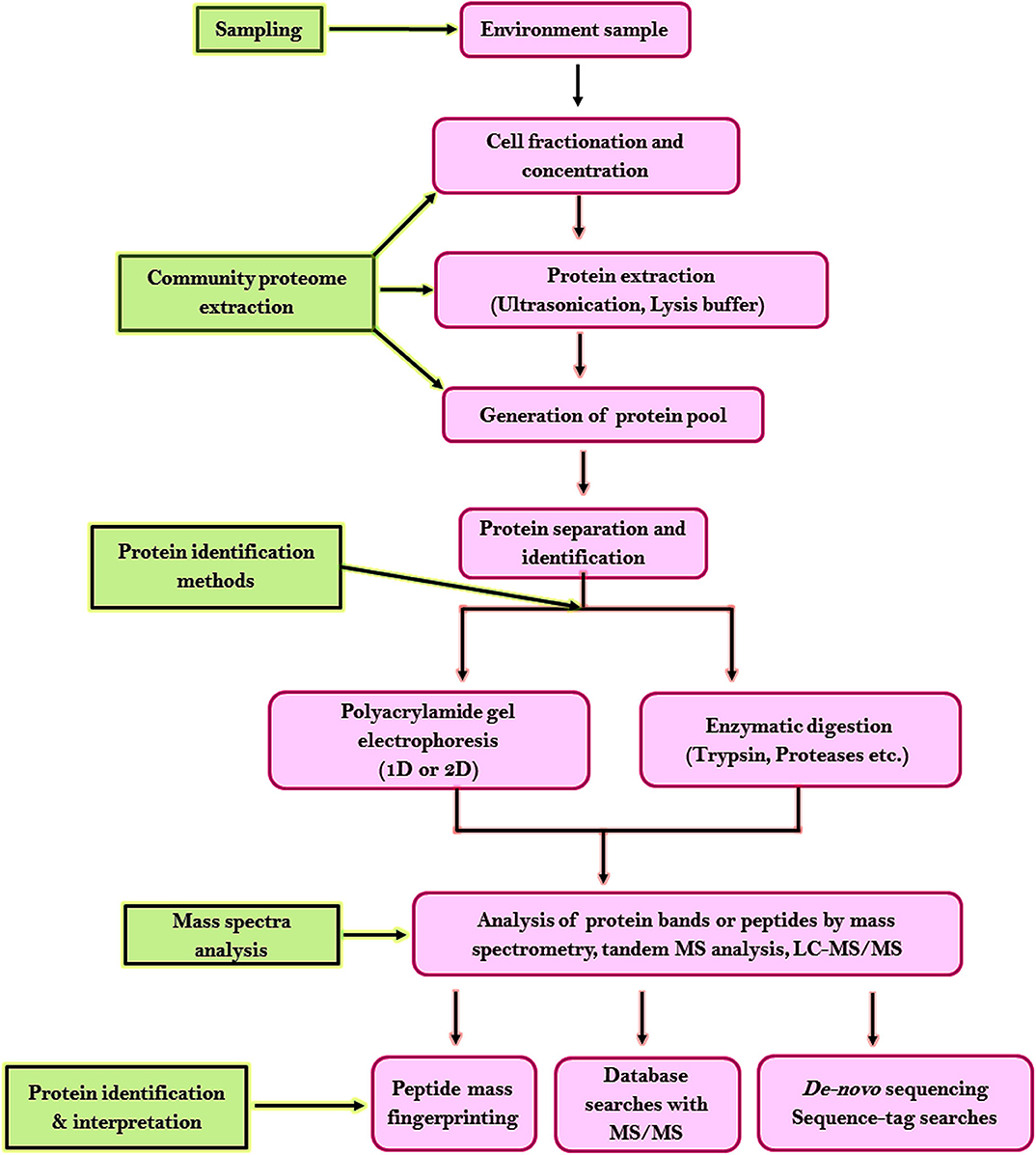
Figure 6 . Workflow for proteomic analysis of microbial community studies.
For comparative and quantitative proteomics data with high reproducibility and precision is useful. It aims to determine the divergence in protein expression in various environmental gradients (e.g., nutrient and salinity gradients) as well as biological conditions like control vs. treatment, healthy vs. disease, specific genotype vs. wild type. Various labeling techniques have been developed for proteomics like stable isotope labeling using amino acids, peptides ( Geiger et al., 2011 ; Lau et al., 2014 ), tandem mass tags ( Ward et al., 2014 ), isotope dilution ( Bettmer, 2010 ), isotope-coded affinity tags ( Hägglund et al., 2008 ), isobaric tags for relative and absolute quantification ( Martínez-Esteso et al., 2014 ). A label-free quantitative proteomic approach is possible with the progress in computational software which allows extensive profiling of proteins with flexibility. The label-free method is deployed on the abundance of proteins by comparing the peak intensity or the spectral count of the same peptide/protein. The mass-spectrometry based label-free approach is cost-effective, more prevalent, and has become the key research methodology in metaproteomics.
The metabolism of an organism is dependent on environmental factors. Changing external stimulus triggers changes in protein expression and assessing such changes can be effective in bioremediation approaches ( Mattarozzi et al., 2017 ). Several studies report the characterization and quantification of proteins expressed by microorganisms present in diverse ecosystems like soil, marine, and freshwater environments, sediment, soil, activated sludge, acid mine drainage biofilms, human or animal microbiome and plant-associated microorganisms ( Siggins et al., 2012 ; Wang et al., 2016 ). Metaproteome data from such ecosystems helps in understanding microbial community structure, dynamics, and functioning. Metaproteomic analysis together with community genomics has been reported to be a dynamic tool to discover microbial diversity and to differentiate closely related organisms in a biofilm colonizing acid mine drainage ( Simmons et al., 2008 ; Denef et al., 2009 , 2010 ). Metaproteomics was reportedly used to analyze an anaerobic microbial community degrading toluene ( Jehmlich et al., 2010 ). Siggins et al. (2012) investigated the impact of temperature and exposure to trichloroethylene (TCE) on proteins expression by the microbial community in laboratory-scale anaerobic conditions.
Metaproteomics has also been reportedly used to distinguish metabolic activities of microbes significant to the bioremediation of contaminated environments. The expression of catabolic enzymes, such as catechol 2,3-dioxygenases, 1,2-cis-dihydrodiol dehydrogenase, and 2-hydroxymuconic semialdehyde was reported in Sphingomonadales and uncultured bacteria assisting the bioremediation of compost ( Bastida et al., 2016 ). A culture-dependent community proteomic study reported that soil microbial community becomes convoluted in hydrocarbon-contaminated soil compared to untreated soil ( Bastida et al., 2014 ).
It can be utilized to explain complex community interplay associated with in situ bioremediation of contaminated soil sites ( Bastida et al., 2010 ). Morris et al. (2010) conducted a comparative metaproteomic study to investigate the membrane proteins expressed by the microbial communities from surface waters in the South Atlantic. A combination of metaproteomics with geochemical and biological data was used to examine the behavior of biofilm-forming microbial communities along environmental gradients. Jehmlich et al. (2010) reported the metaproteome of an anaerobic community, initiating from a batch sulfate reducing enrichment culture, exposed to toluene. A metaproteome analysis in bioremediation thus encompasses the identification of differentially expressed proteins and their respective genes, assessing changes in the abundance of biomarkers, protein structure, and function characterization.
The metabolic adaptation of Arthrobacter phenanthrenivorans Sphe3 supplemented with phenanthrene, phthalate, glucose as sole carbon and energy source was investigated using comparative quantitative proteomics. A gel-free quantitative proteomic analysis using nano LC-MS/MS was used to explore the phenanthrene catabolic pathway as well as the fluctuations in proteins when A. phenanthrenivorans Sphe3 cells were grown on various substrates ( Vandera et al., 2015 ). A proteomic study using nano LC-MS/MS has contributed awareness into the metabolic and regulatory networks of Psuedomonas aeruginosa san ai exposed to plastic additive 2,6-di-tert-butylphenol ( Medić et al., 2019 ). LC–MS/MS shotgun proteomics was reportedly used to determine variations in the proteome of hydrocarbon-degrading psychrophile Oleispira antarctica RB-8 when grown on n -alkanes in cold temperatures ( Gregson et al., 2020 ).
Metabolomics
A metabolome is the total metabolites in an organism and the study of the metabolite profile of a cell within a given condition is called metabolomics ( Beale et al., 2017 ). A cell produces a range of primary and secondary metabolites when subjected to external stress which has enabled us to understand and analyze the effect of environmental conditions on the metabolome of organisms ( Malla et al., 2018 ). Metabolome based approaches have facilitated to establish models that can predict microbial activities under bioremediation strategies. It permits us to comprehend the potent activities of the microbial communities and their functional inputs to the environments in which they flourish. Various spectroscopic techniques can be used to characterize metabolites produced by organisms ( Bargiela et al., 2015 ).
Metabolomics analyzes the metabolites produced by the cell in response to changing environmental conditions which in turn provide information about the regulatory events in a cell ( Krumsiek et al., 2015 ). These metabolites can be used as bioindicators to screen the biological effects of the contaminants waste for a better perception of the environment. Advances in analytical instruments, statistical tools and bioinformatics tools made access to assess, extract, and interpret various metabolites and elucidate their pathway ( Hill et al., 2015 ). The major strategies to analyze the metabolome involve metabolite profiling, targeted analysis, and metabolic fingerprinting ( Wang et al., 2010 ). Metabolite profiling is a semi-quantitative process that enables the detection of both known and unknown metabolites while target analysis is a quantitative approach that uses various detection techniques to detect metabolites involved in a specific pathway. Metabolic fingerprinting is a high throughput procedure that generates fingerprints of specific metabolites in a sample.
A metabolomics analysis workflow starts with sample acquisition and preparation followed by separation and detection of analytes. Detection and quantification of metabolites is normally accomplished through an amalgamation of chromatography techniques (liquid chromatography and gas chromatography) and detection systems like mass spectrometry and nuclear magnetic resonance ( Aldridge and Rhee, 2014 ). The most extensively deployed analytical instruments in metabolomics research are nuclear magnetic resonance spectrometers and mass spectrometry ( Healy et al., 1995 ; Pan and Raftery, 2007 ). Among the mass spectrometry (MS) technology liquid chromatography MS is widely used due to lesser ion suppression, higher resolution, and non-requirement of chemical derivatization steps. Mass spectrometry produces spectra comprising of a pattern of peaks that can be used to analyze and quantify metabolites. These patterns are saved in spectral databases, allowing automated analysis, and generation of metabolomic profiles. Two methods are commonly used to analyze metabolic data. The first method uses statistics and clustering algorithms while the second method makes use of networks to envisage the spatial and temporal properties of the data. Several software tools like MetaboAnalyst, XCMS, MetATT, Matlab, etc. are used for metabolomics data analysis. Apart from data analysis several network visualization software tools like MetScape Plugin, MetaMapp, MAVEN, Pathomx, etc. have been developed to visualize graphical representations of metabolic pathways as diagrams or heat maps ( Hill et al., 2015 ).
Mallick et al. (2019) used a computational framework called Model-based Genomically Informed High-dimensional Predictor of Microbial Community Metabolic Profiles (MelonnPan) to predict community metabolomes from a microbial community. This approach predicted and correlated the possible metabolic profiles across many samples at low cost compared to metabolomics, thus offering new opportunities for economical study designs. It also enabled the identification of functionally pertinent gene families with species or strain-specific metabolic associations, expediting biologically significant mechanisms at finer taxonomic resolution. This analysis in the future may thus provide metabolic understandings and hypothesis generation in numerous samples for which only metagenomic data are currently available.
Metabolomics along with bioinformatics tools and databases have enabled a better understanding of microbial community, their catabolic pathways, and genes responsible for encoding catabolic enzymes. It is thus a powerful strategy for discovering novel metabolic pathways and characterizing metabolic networks ( Villas-Bôas et al., 2007 ; Kim et al., 2016 ; McMahon et al., 2018 ). Metagenomic, geochemical, and metabolomic analyses were used to study the role of microbial communities in the degradation of detrital biomass and petroleum hydrocarbons ( Keum et al., 2008 ; Dong et al., 2019 ). Targeted and untargeted metabolomics using LC/MS/MS system was used to investigate reprogrammed-metabolism underlying biofilm formation for deciphering the metabolism for biofilm formation ( Lu et al., 2019 ). Proteogenomic and metabolomic approaches were used to identify the pathways and enzymes used by marine bacteria Mycobacterium sp. DBP42 and Halomonas sp. ATBC28 to degrade plasticizers like dibutyl phthalate (DBP), bis(2-ethyl hexyl) phthalate (DEHP), and acetyl tributyl citrate (ATBC) ( Wright et al., 2020 ).
Metabolite pathway databases and repositories can be used to supervise, investigate, and disseminate information about metabolites and their pathways. They provide a databank on metabolic information as well as helps in the unification of complex data into metabolic pathways. These databases and repositories also help in modeling metabolic pathways that can be investigated and prompted using mathematical modeling techniques. A variety of pathway databases and repositories, such as Reactome, KEGG (Kyoto Encyclopedia of Genes and Genomes), PANTHER pathway, etc. are available which provide information about specific metabolite pathways, reactions and enzymes, names, chemical structures, molecular weight, the occurrence of metabolites in pathways, etc. ( Hill et al., 2015 ). Metabolomics is thus promptly expanding omic technology that can provide integrated insight into various metabolites and metabolic pathways. The collective uses of modern instrumental analytical methodologies, bioinformatics softwares have unraveled the detection of metabolites that cannot be accomplished by single-analysis techniques. Advancement in analytical platforms will boost the incorporation of metabolomics into systems biology.
Fluxomics is a quantitative approach that studies the rates of metabolic reactions, changes in metabolic rates inside a biological individual. Fluxome is the complete set of metabolic fluxes in a cell which provides facts on several cellular processes, thus it is a distinctive phenotypic characteristic of cells. Flux analysis offers vital information about the phenotype since it assesses the metabolome in its functional communications with the genome and environment ( Dettmer et al., 2007 ). Fluxomic approaches analyze metabolic interfaces prominent to the progression and logical design of cellular functions. Metabolic reaction rates evaluate the in vivo enzymatic activities that cannot be obtained from the transcriptomic, proteomic, or metabolomic data alone. A successful fluxomic analysis is dependent on the precision of quantitative metabolomics data, isotopic labeling measurements, and the reorganization of metabolic networks describing stoichiometry and regulation of metabolic reactions.
Two techniques commonly used for flux quantification are flux balance analysis or stoichiometric metabolic flux analysis and tracer-based metabolomics which uses a firm isotope of carbon 13 C to trace the carbon distributing through diverse pathways followed by analytical techniques like nuclear magnetic resonance (NMR) or mass spectroscopy (MS) analysis for identification and characterization of the labeled compounds. Flux balance analysis uses the stoichiometry of metabolic reactions in concurrence with biological, chemical, and thermodynamic parameters to produce a controlled model of metabolic flux. In 13 C isotope labeling, cells are fed with 13 C labeled substrates which can be metabolized (e.g., Glucose) resulting in metabolites containing 13 C atoms. Depending on the metabolic pathway 13 C atoms are amalgamated into the newly designed metabolites in diverse positions and numbers. Therefore, for every single metabolite, there may be various isotopomers (numerous molecules of the same metabolite with different labeling states). Metabolomic analytical platforms are used to regulate the concentration and isotopomeric dissemination of these metabolites ( Lee and Go, 2005 ).
The comparative and analytical approaches have been used as valid methods to estimate flux ratios. Tracer-based metabolomics data can be used to analyze metabolic flux distribution by using the comparative analytical approach and the integrative approach. Comparative analysis of tracer-based metabolomics data is also known as MIDA (mass isotopomer distribution analysis) which does not require bioinformatics tools and requires minimal statistics knowledge. It is a functional tool for the characterization of cell metabolic flux distribution by comparing the labeling patterns obtained with appropriate labeling distributions depending on the metabolic pathway followed by the tracer ( Hellerstein and Neese, 1999 ; Kumari et al., 2018 ; Barupal et al., 2020 ). It provides predictable estimates of relative metabolic flux distributions. The perceptive nature of this approach has contributed to its extensive use in the interpretation of tracer-based experiments. This approach suffers the limitation that it does not allow the full characterization of the fluxome.
The integrative approach uses the integration of tracer-based metabolomic data into software packages to estimate the flux distribution by fitting the experimental data into the framework of more complete metabolic models ( Wiechert, 2001 ). The 13 C-based metabolic flux investigation has been used mainly to quantify the intracellular fluxes of an extensive range of microorganisms ( Sauer, 2006 ). Software packages like 13 C-FLUX have been developed which calculate sets of fluxes ( Wiechert et al., 2001 ). The fluxome profile evaluation of marine bacteria Shewanella using biochemical, GC-MS analyzes, statistical and genetic algorithms revealed the presence of co-metabolic routes for the biodegradation of lethal metals, halogenated organic compounds, and radionuclides. The bacteria exhibited a responsive metabolic flux when revealed toward different carbon sources ( Tang et al., 2007 ). Consolidated molecular diagnostic and kinetic modeling enhanced cognizance about organohalide-respiring bacteria (ORB) and their metabolite dynamics in an in-situ dechlorinating bioreactor ( Maphosa et al., 2010b ). Metabolic flux investigation revealed valuable evidence of a phenol overproducing strain resulting from Psuedomonas putida S12 during metabolic engineering ( Wierckx et al., 2008 , 2009 ). Metabolic flux analysis was used to investigate the constitutive metabolic network for the co-utilization of sugar and aromatic carbons in Pseudomonas putida ( Kukurugya et al., 2019 ). Flux distributions using 13 C-MFA (Metabolic flux analysis) was used to identify the effect of phenol on the carbon metabolism in wild type E. coli cultured under varying phenol concentrations ( Kitamura et al., 2019 ).
Flux analysis can provide strategies for optimizing a biological system. Fluxomics is expected to progress and provide significant novel results for the development of biological systems and systems biology. Technical difficulties in the development of flux analysis and intricate metabolic regulatory mechanisms have inadequate the opportunity of fluxomics. It is thus essential to combine high throughput “omics” and molecular biology techniques to insistence and clarify challenges in the fluxomics for its future development.
The immense threat posed to the environment due to anthropogenic activities has led to think about novel strategies for decontamination and clean up. It is challenging to realize and discover the interplay between the microbial communities in polluted environments. Modern era omic-approaches like genomics, proteomics, transcriptomics, metabolomics, and fluxomics have eliminated the boundaries to study the mechanisms involved in various bioremediation pathways. It has enabled to incorporate new strategies for efficient bioremediation process. Omics-approaches have the potency to anticipate microbial metabolism in polluted environments. These high-throughput analyses would aid in tracking novel organisms for bioremediation, provide excellent and novel insights into their key biodegradative pathways at the molecular level. These multi-omics approaches will enable us to make new hypotheses, theories, and archetypes for bioremediation of polluted surroundings.
Author Contributions
MM and HC conceived the idea of the review, provided the general concept and inputs for each specific section, drafted part of the manuscript, and wrote the review after collecting literature. MM edited, compiled, and finalized the draft. HC, MM, and KS read and approved it for publication. All authors contributed to the article and approved the submitted version.
Conflict of Interest
The authors declare that the research was conducted in the absence of any commercial or financial relationships that could be construed as a potential conflict of interest.
Acknowledgments
MM was thankful to Mohanlal Sukhadia University, Udaipur for providing the necessary facilities during the course of study. This study was supported by Startup Research Grant (UGC Faculty Research Promotion Scheme; FRPS) and sustained by Mohanlal Sukhadia University, Udaipur, Rajasthan, India.
Abou Seeda, M. A., Yassen, A. A., and Abou El-Nour, E. Z. A. A. (2017). Microorganism as a tool of bioremediation technology for cleaning waste and industrial water. Biosci. Res. 14, 633–636.
Adessi, C., Matton, G., Ayala, G., Turcatti, G., Mermod, J. J., Mayer, P., et al. (2000). Solid phase DNA amplification: characterisation of primer attachment and amplification mechanisms. Nucleic Acids Res . 28:E87. doi: 10.1093/nar/28.20.e87
PubMed Abstract | CrossRef Full Text | Google Scholar
Aldridge, B. B., and Rhee, K. Y. (2014). Microbial metabolomics: innovation, application, insight. Curr. Opin. Microbiol. 19, 90–96. doi: 10.1016/j.mib.2014.06.009
Arazoe, T., Kondo, A., and Nishida, K. (2018). Targeted nucleotide editing technologies for microbial metabolic engineering. Biotechnol. J. 13:e1700596. doi: 10.1002/biot.201700596
Aubé, J., Senin, P., Bonin, P., Pringault, O., Jeziorski, C., Bouchez, O., et al. (2020). Meta-omics provides insights into the impact of hydrocarbon contamination on microbial mat functioning. Microb. Ecol. 80, 286–295. doi: 10.1007/s00248-020-01493-x
Awasthi, M. K., Ravindran, B., Sarsaiya, S., Chen, H., Wainaina, S., Singh, E., et al. (2020). Metagenomics for taxonomy profiling: tools and approaches. Bioengineered 11, 356–374. doi: 10.1080/21655979.2020.1736238
PubMed Abstract | CrossRef Full Text
Bahamonde, P. A., Feswick, A., Isaacs, M. A., Munkittrick, K. R., and Martyniuk, C. J. (2016). Defining the role of omics in assessing ecosystem health: perspectives from the Canadian environmental monitoring program. Environ. Toxicol. Chem . 35, 20–35. doi: 10.1002/etc.3218
Bargiela, R., Herbst, F. A., Martínez-Martínez, M., Seifert, J., Rojo, D., Cappello, S., et al. (2015). Metaproteomics and metabolomics analyses of chronically petroleum-polluted sites reveal the importance of general anaerobic processes uncoupled with degradation. Proteomics 15, 3508–3520. doi: 10.1002/pmic.201400614
Barupal, T., Chittora, D., and Meena, M. (2019a). “Solid waste: characterization, assessment, monitoring, and remediation,” in Advances in Waste-to-Energy Technologies , eds R. P. Singh, V. Prasad, and B. Vaish (Boca Raton; London; New York, NY: Taylor & Francis; CRC Press), 1–19. doi: 10.1201/9780429423376-1
CrossRef Full Text
Barupal, T., Meena, M., and Sharma, K. (2019b). Inhibitory effects of leaf extract of Lawsonia inermis on Curvularia lunata and characterization of novel inhibitory compounds by GC–MS analysis. Biotechnol. Rep. 23:e00335. doi: 10.1016/j.btre.2019.e00335
Barupal, T., Meena, M., and Sharma, K. (2020). A study on preventive effects of Lawsonia inermis L. bioformulations against leaf spot disease of maize. Biocatal. Agric. Biotechnol. 23:101473. doi: 10.1016/j.bcab.2019.101473
CrossRef Full Text | Google Scholar
Bastida, F., Hernandez, T., and Garcia, C. (2014). Metaproteomics of soils from semiarid environment: functional and phylogenetic information obtained with different protein extraction methods. J. Proteomics 101, 31–42. doi: 10.1016/j.jprot.2014.02.006
Bastida, F., Jehmlich, N., Lima, K., Morris, B. E. L., Richnow, H. H., Hernández, T., et al. (2016). The ecological and physiological responses of the microbial community from a semiarid soil to hydrocarbon contamination and its bioremediation using compost amendment. J. Proteomics 135, 162–169. doi: 10.1016/j.jprot.2015.07.023
Bastida, F., Nicolás, C., Moreno, J., Hernández, T., and Garcia, C. (2010). Tracing changes in the microbial community of a hydrocarbon-polluted soil by culture-dependent proteomics. Pedosphere 20, 479–485. doi: 10.1016/S1002-0160(10)60037-9
Beale, D. J., Karpe, A. V., Ahmed, W., Cook, S., Morrison, P. D., Staley, C., et al. (2017). A community multi-omics approach towards the assessment of surface water quality in an urban river system. Int. J. Environ. Res. Public Health 14:E303. doi: 10.3390/ijerph14030303
Bentley, D. R., Balasubramanian, S., Swerdlow, H. P., Smith, G. P., Milton, J., Brown, C. B., et al. (2008). Accurate whole human genome sequencing using reversible terminator chemistry. Nature 456, 53–59. doi: 10.1038/nature07517
Bettmer, J. (2010). Application of isotope dilution ICP-MS techniques to quantitative proteomics. Anal. Bioanal. Chem. 397, 3495–3502. doi: 10.1007/s00216-010-3861-y
Bier, E., Harrison, M. M., O'Connor-Giles, K. M., and Wildonger, J. (2018). Advances in engineering the fly genome with the CRISPR-Cas system. Genetics 208, 1–18. doi: 10.1534/genetics.117.1113
Bilal, T., Malik, B., and Hakeem, K. R. (2018). Metagenomic analysis of uncultured microorganisms and their enzymatic attributes. J. Microbiol. Methods 155, 65–69. doi: 10.1016/j.mimet.2018.11.014
Braslavsky, I., Hebert, B., Kartalov, E., and Quake, S. R. (2003). Sequence information can be obtained from single DNA molecules. Proc. Natl. Acad. Sci. U.S.A. 100, 3960–3964. doi: 10.1073/pnas.0230489100
Bursle, E., and Robson, J. (2016). Non-culture methods for detecting infection. Aust. Prescr. 39, 171–175. doi: 10.18773/austprescr.2016.059
Callaghan, A. V. (2013). Metabolomic investigations of anaerobic hydrocarbon-impacted environments. Curr. Opin. Biotechnol. 24, 506–515. doi: 10.1016/j.copbio.2012.08.012
Cárdenas Espinosa, M. J., Colina Blanco, A., Schmidgall, T., Atanasoff-Kardjalieff, A. K., Kappelmeyer, U., Tischler, D., et al. (2020). Toward biorecycling: isolation of a soil bacterium that grows on a polyurethane oligomer and monomer. Front. Microbiol. 11:404. doi: 10.3389/fmicb.2020.00404
Chakraborty, R., Wu, C. H., and Hazen, T. C. (2012). Systems biology approach to bioremediation. Curr. Opin. Biotechnol. 23, 483–490. doi: 10.1016/j.copbio.2012.01.015
Chandran, H., Meena, M., Barupal, T., and Sharma, K. (2020). Plant tissue culture as a perpetual source for production of industrially important bioactive compounds. Biotechnol. Rep. 26:e00450. doi: 10.1016/j.btre.2020.e00450
Chen, Y., and Murrell, J. C. (2010). When metagenomics meets stable-isotope probing: progress and perspectives. Trends Microbiol. 18, 157–163. doi: 10.1016/j.tim.2010.02.002
Chittora, D., Meena, M., Barupal, T., Swapnil, P., and Sharma, K. (2020). Cyanobacteria as a source of biofertilizers for sustainable agriculture. Biochem. Biophys. Rep. 22:100737. doi: 10.1016/j.bbrep.2020.100737
Cho, J. C., and Tiedje, J. M. (2001). Bacterial species determination from DNA-DNA hybridization by using genome fragments and DNA microarrays. Appl. Environ. Microbiol. 67, 3677–3682. doi: 10.1128/AEM.67.8.3677-3682.2001
Cho, J. C., and Tiedje, J. M. (2002). Quantitative detection of microbial genes by using DNA microarrays. Appl. Environ. Microbiol. 68, 1425–1430. doi: 10.1128/AEM.68.3.1425-1430.2002
Clarridge, J. E. (2004). Impact of 16S rRNA gene sequence analysis for identification of bacteria on clinical microbiology and infectious diseases. Clin. Microbiol. Rev. 17, 840–862. doi: 10.1128/CMR.17.4.840-862.2004
Dai, Z., Zhang, S., Yang, Q., Zhang, W., Qian, X., Dong, W., et al. (2018). Genetic tool development and systemic regulation in biosynthetic technology. Biotechnol. Biofuels 11:152. doi: 10.1186/s13068-018-1153-5
Daniel, R. (2005). The metagenomics of soil. Nat. Rev. Microbiol. 3, 470–478. doi: 10.1038/nrmicro1160
Das, D., Mawlong, G. T., Sarki, Y. N., Singh, A. K., Chikkaputtaiah, C., and Boruah, H. P. D. (2020). Transcriptome analysis of crude oil degrading Pseudomonas aeruginosa strains for identification of potential genes involved in crude oil degradation. Gene 755:144909. doi: 10.1016/j.gene.2020.144909
De Sousa, C. S., Hassan, S. S., Pinto, A. C., Silva, W. M., De Almeida, S. S., Soares, S. D. C., et al. (2018). “Microbial omics: applications in biotechnology,” in Omics Technologies and Bio-Engineering , eds D. Barh and V. Azevedo (Cambridge, MA: Academic Press), 3–20. doi: 10.1016/B978-0-12-815870-8.00001-2
Denef, V. J., Kalnejais, L. H., Mueller, R. S., Wilmes, P., Baker, B. J., Thomas, B. C., et al. (2010). Proteogenomic basis for ecological divergence of closely related bacteria in natural acidophilic microbial communities. Proc. Natl. Acad. Sci. U.S.A. 107, 2383–2390. doi: 10.1073/pnas.0907041107
Denef, V. J., VerBerkmoes, N. C., Shah, M. B., Abraham, P. E., Lefsrud, M., Hettich, R. L., et al. (2009). Proteomics-inferred genome typing (PIGT) demonstrates inter-population recombination as a strategy for environmental adaptation. Environ. Microbiol. 11, 313–325. doi: 10.1111/j.1462-2920.2008.01769.x
Dennis, P., Edwards, E. A., Liss, S. N., and Fulthorpe, R. (2003). Monitoring gene expression in mixed microbial communities by using DNA microarrays. Appl. Environ. Microbiol . 69, 769–778. doi: 10.1128/AEM.69.2.769-778.2003
Desai, C., Pathak, H., and Madamwar, D. (2010). Advances in molecular and ‘-omics’ technologies to gauge microbial communities and bioremediation at xenobiotic/anthropogen contaminated sites. Bioresour. Technol. 101, 1558–1569. doi: 10.1016/j.biortech.2009.10.080
Dettmer, K., Aronov, P. A., and Hammock, B. D. (2007). Mass spectrometry based metabolomics. Mass Spectrom. Rev. 26, 51–58. doi: 10.1002/mas.20108
Dickson, R. P., Erb-Downward, J. R., Prescott, H. C., Martinez, F. J., Curtis, J. L., Lama, V. N., et al. (2014). Analysis of culture-dependent versus culture-independent techniques for identification of bacteria in clinically obtained bronchoalveolar lavage fluid. J. Clin. Microbiol. 52, 3605–3613. doi: 10.1128/JCM.01028-14
Dietrich, J. A., McKee, A. E., and Keasling, J. D. (2010). High-throughput metabolic engineering: advances in small-molecule screening and selection. Annu. Rev. Biochem. 79, 563–590. doi: 10.1146/annurev-biochem-062608-095938
Dinsdale, E. A., Edwards, R. A., Hall, D., Angly, F., Breitbart, M., Brulc, J. M., et al. (2008). Functional metagenomic profiling of nine biomes. Nature 452, 629–632. doi: 10.1038/nature06810
Dong, X., Greening, C., Rattray, J. E., Chakraborty, A., Chuvochina, M., Mayumi, D., et al. (2019). Metabolic potential of uncultured bacteria and archaea associated with petroleum seepage in deep-sea sediments. Nat. Commun. 10:1816. doi: 10.1038/s41467-019-09747-0
Eid, J., Fehr, A., Gray, J., Luong, K., Lyle, J., Otto, G., et al. (2009). Real-time DNA sequencing from single polymerase molecules. Science 323, 133–138. doi: 10.1126/science.1162986
El-Metwally, S., Ouda, O. M., and Helmy, M. (2014). Next Generation Sequencing Technologies and Challenges in Sequence Assembly . New York, NY: Springer Verlag.
Eyers, L., George, I., Schuler, L., Stenuit, B., Agathos, S. N., and El Fantroussi, S. (2004). Environmental genomics: exploring the unmined richness of microbes to degrade xenobiotics. Appl. Microbiol. Biotechnol. 66, 123–130. doi: 10.1007/s00253-004-1703-6
Fedurco, M., Romieu, A., Williams, S., Lawrence, I., and Turcatti, G. (2006). BTA, a novel reagent for DNA attachment on glass and efficient generation of solid-phase amplified DNA colonies. Nucleic Acids Res. 34:e22. doi: 10.1093/nar/gnj023
Felczykowska, A., Krajewska, A., Zielińska, S., Łoś, J. M., Bloch, S. K., and Nejman-Faleńczyk, B. (2015). The most widespread problems in the function-based microbial metagenomics. Acta Biochim. Pol. 62, 161–166. doi: 10.18388/abp.2014_917
Ferrer, M., Beloqui, A., Vieites, J. M., Guazzaroni, M. E., Berger, I., and Aharoni, A. (2009). Interplay of metagenomics and in vitro compartmentalization. Microb. Biotechnol. 2, 31–39. doi: 10.1111/j.1751-7915.2008.00057.x
Gao, H., Wang, Y., Liu, X., Yan, T., Wu, L., Alm, E., et al. (2004). Global transcriptome analysis of the heat shock response of Shewanella oneidensis . J. Bacteriol. 186, 7796–7803. doi: 10.1128/JB.186.22.7796-7803.2004
Garrido-Sanz, D., Redondo-Nieto, M., Guirado, M., Pindado Jiménez, O., Millán, R., Martin, M., et al. (2019). Metagenomic insights into the bacterial functions of a diesel-degrading consortium for the rhizoremediation of diesel-polluted soil. Genes 10:456. doi: 10.3390/genes10060456
Gaytán, I., Sánchez-Reyes, A., Burelo, M., Vargas-Suárez, M., Liachko, I., and Press, M. (2020). Degradation of recalcitrant polyurethane and xenobiotic additives by a selected landfill microbial community and its biodegradative potential revealed by proximity ligation-based metagenomic analysis. Front. Microbiol. 10:2986. doi: 10.3389/fmicb.2019.02986
Geiger, T., Wisniewski, J. R., Cox, J., Zanivan, S., Kruger, M., Ishihama, Y., et al. (2011). Use of stable isotope labeling by amino acids in cell culture as a spike-in standard in quantitative proteomics. Nat. Protoc. 6, 147–157. doi: 10.1038/nprot.2010.192
Ghazanfar, S., Azim, A., Ghazanfar, M. A., Anjum, M. I., and Begum, I. (2010). Metagenomics and its application in soil microbial community studies: biotechnological prospects. J. Anim. Plant Sci. 6, 611–622.
Google Scholar
Gilbert, J. A., and Dupont, C. L. (2011). Microbial metagenomics: beyond the genome. Annu. Rev. Mar. Sci. 3, 347–371. doi: 10.1146/annurev-marine-120709-142811
Godzik, A. (2011). Metagenomics and the protein universe. Curr. Opin. Struct. Biol. 21, 398–403. doi: 10.1016/j.sbi.2011.03.010
Gołebiewski, M., and Tretyn, A. (2020). Generating amplicon reads for microbial community assessment with next generation sequencing. J. Appl. Microbiol . 128, 330–354. doi: 10.1111/jam.14380
Greene, E. A., and Voordouw, G. (2003). Analysis of environmental microbial communities by reverse sample genome probing. J. Microbiol. Methods 53, 211–219. doi: 10.1016/S0167-7012(03)00024-1
Gregson, B. H., Metodieva, G., Metodiev, M. V., Golyshin, P. N., and McKew, B. A. (2020). Protein expression in the obligate hydrocarbon-degrading psychrophile Oleispira antarctica RB-8 during alkane degradation and cold tolerance. Environ. Microbiol. 22, 1870–1883. doi: 10.1111/1462-2920.14956
Gupta, K., Biswas, R., and Sarkar, A. (2020). “Advancement of omics: prospects for bioremediation of contaminated soils,” in Microbial Bioremediation & Biodegradation , ed M. Shah (Singapore: Springer), 113–42.
Gutierrez, D. B., Gant-Branum, R. L., Romer, C. E., Farrow, M. A., Allen, J. L., Dahal, N., et al. (2018). An integrated, high-throughput strategy for “multi-omic” systems level analysis. J. Proteome Res. 17, 3396–3408. doi: 10.1021/acs.jproteome.8b00302
Gutleben, J., Chaib De Mares, M., van Elsas, J. D., Smidt, H., Overmann, J., and Sipkema, D. (2018). The multi-omics promise in context: From sequence to microbial isolate. Crit. Rev. Microbiol . 44, 212–229. doi: 10.1080/1040841X.2017.1332003
Hägglund, P., Bunkenborg, J., Maeda, K., and Svensson, B. (2008). Identification of thioredoxin disulfide targets using a quantitative proteomics approach based on isotope-coded affinity tags. J. Proteome Res. 7, 5270–5276. doi: 10.1021/pr800633y
Hallin, P. F., Binnewies, T. T., and Ussery, D. W. (2008). The genome BLAST atlas-a Gene Wiz extension for visualization of whole-genome homology. Mol. Biosyst. 4, 363–371. doi: 10.1039/b717118h
Han, D., Gao, P., Li, R., Tan, P., Xie, J., Zhang, R., et al. (2020). Multicenter assessment of microbial community profiling using 16S rRNA gene sequencing and shotgun metagenomic sequencing. J. Adv. Res. doi: 10.1016/j.jare.2020.07.010
Handelsman, J. (2004). Metagenomics: application of genomics to uncultured microorganisms. Microbiol. Mol. Bio. Rev. 68, 669–685. doi: 10.1128/MMBR.68.4.669-685.2004
Hart, E. H., Creevey, C. J., Hitch, T., and Kingston-Smith, A. H. (2018). Meta-proteomics of rumen microbiota indicates niche compartmentalisation and functional dominance in a limited number of metabolic pathways between abundant bacteria. Sci. Rep. 8:10504. doi: 10.1038/s41598-018-28827-7
He, Z., Deng, Y., van Nostrand, J. D., Xu, M., Hemme, L. H., Tu, Q., et al. (2010). GeoChip 3.0 as a high-throughput tool for analyzing microbial community composition, structure and functional activity. ISME J. 4, 1167–1179. doi: 10.1038/ismej.2010.46
He, Z., Zhang, P., Wu, L., Rocha, A. M., Tu, Q., Shi, Z., et al. (2018). Microbial functional gene diversity predicts groundwater contamination and ecosystem functioning. mBio. 9:e02435-17. doi: 10.1128/mBio.02435-17
Head, I. M., Singleton, I., and Milner, M. G. (2003). Bioremediation: A Critical Review . Wymondham: Horizon Scientific Press.
Healy, F. G., Ray, R. M., Aldrich, H. C., Wilkie, A. C., Ingram, L. O., and Shanmugam, K. T. (1995). Direct isolation of functional genes encoding cellulases from the microbial consortia in a thermophilic, anaerobic digester maintained on lignocelluloses. Appl. Microbiol. Biotechnol. 43, 667–674. doi: 10.1007/BF00164771
Hellerstein, M. K., and Neese, R. A. (1999). Mass isotopomer distribution analysis at eight years: theoretical, analytic, and experimental considerations. Am. J. Physiol. 276, E1146–E1170. doi: 10.1152/ajpendo.1999.276.6.E1146
Hill, C. B., Czauderna, T., Klapperstück, M., Roessner, U., and Schreiber, F. (2015). Metabolomics, standards, and metabolic modeling for synthetic biology in plants. Front. Bioeng. Biotechnol. 3:167. doi: 10.3389/fbioe.2015.00167
Hillmann, B., Al-Ghalith, G. A., Shields-Cutler, R. R., Zhu, Q., Gohl, D. M., Beckman, K. B., et al. (2018). Evaluating the information content of shallow shotgun metagenomics. mSystems 3:e00069-18. doi: 10.1128/mSystems.00069-18
Hodkinson, B. P., and Grice, E. A. (2015). Next-generation sequencing: a review of technologies and tools for wound microbiome research. Adv. Wound Care 4, 50–58. doi: 10.1089/wound.2014.0542
Igiri, B. E., Okoduwa, S. I. R., Idoko, G. O., Akabuogu, E. P., Adeyi, A. O., and Ejiogu, I. K. (2018). Toxicity and bioremediation of heavy metals contaminated ecosystem from tannery wastewater: a review. J. Toxicol. 2018:2568038. doi: 10.1155/2018/2568038
Iwamoto, T., and Nasu, M. (2001). Current bioremediation practice and perspective. J. Biosci. Bioeng. 92, 1–8. doi: 10.1016/S1389-1723(01)80190-0
Jain, M., Hugh, E. O., Paten, B., and Akeson, M. (2016). The oxford nanopore MinION: Delivery of nanopore sequencing to the genomics community. Genome Biol. 17:239. doi: 10.1186/s13059-016-1103-0
Jeffries, T. C., Rayu, S., Nielsen, U. N., Lai, K., Ijaz, A., Nazaries, L., et al. (2018). Metagenomic functional potential predicts degradation rates of a model organophosphorus xenobiotic in pesticide contaminated soils. Front. Microbiol. 9:147. doi: 10.3389/fmicb.2018.00147
Jehmlich, N., Kleinsteuber, S., Vogt, C., Benndorf, D., Harms, H., Schmidt, F., et al. (2010). Phylogenetic and proteomic analysis of an anaerobic toluene-degrading community. J. Appl. Microbiol . 109, 1937–1945. doi: 10.1111/j.1365-2672.2010.04823.x
Kapley, A., Prasad, S., and Purohit, H. J. (2007). Changes in microbial diversity in fed-batch reactor operation with wastewater containing nitroaromatic residues. Bioresour. Technol. 98, 2479–2484. doi: 10.1016/j.biortech.2006.09.012
Katsaveli, K., Vayenas, D., Tsiamis, G., and Bourtzis, K. (2012). Bacterial diversity in Cr(VI) and Cr(III)-contaminated industrial wastewaters. Extremophiles 16, 285–296. doi: 10.1007/s00792-012-0429-0
Kchouk, M., Gibrat, J. F., and Elloumi, M. (2017). Generations of sequencing technologies: from first to next generation. Biol. Med. 9:395. doi: 10.4172/0974-8369.1000395
Keum, Y. S., Seo, J. S., Li, Q. X., and Kim, J. H. (2008). Comparative metabolomic analysis of Sinorhizobium sp. C4 during the degradation of phenanthrene. Appl. Microbiol. Biotechnol. 80, 863–872. doi: 10.1007/s00253-008-1581-4
Kim, H., Kwon, K. K., Seong, W., and Lee, S. G. (2016). Multi-enzyme screening using a high-throughput genetic enzyme screening system. J. Vis. Exp. 114:54059. doi: 10.3791/54059
Kim, H. J., Ishidou, E., Kitagawa, E., Momose, Y., and Iwahashi, H. (2004). A yeast DNA microarray for the evaluation of toxicity in environmental water containing burned ash. Environ. Monit. Assess. 92, 253–272. doi: 10.1023/B:EMAS.0000014504.03500.41
Kitamura, S., Toya, Y., and Shimizu, H. (2019). 13 C-Metabolic flux analysis reveals effect of phenol on central carbon metabolism in Escherichia coli . Front. Microbiol. 10:1010. doi: 10.3389/fmicb.2019.01010
Kou, S., Vincent, G., Gonzalez, E., Pitre, F. E., Labrecque, M., and Brereton, N. J. B. (2018). The response of a 16S ribosomal RNA gene fragment amplified community to lead, zinc, and copper pollution in a Shanghai field trial. Front. Microbiol. 9:366. doi: 10.3389/fmicb.2018.00366
Krumsiek, J., Mittelstrass, K., Do, K. T., Stückler, F., Ried, J., Adamski, J., et al. (2015). Gender-specific pathway differences in the human serum metabolome. Metabolomics 11, 1815–1833. doi: 10.1007/s11306-015-0829-0
Kukurugya, M. A., Mendonca, C. M., Solhtalab, M., RWilkes, R. A., Thannhauser, T. W., and Aristilde, L. (2019). Multi-omics analysis unravels a segregated metabolic flux network that tunes co-utilization of sugar and aromatic carbons in Pseudomonas putida . J. Biol. Chem. 294, 8464–8479. doi: 10.1074/jbc.RA119.007885
Kumar, M., and Khanna, S. (2010). Diversity of 16S rRNA and dioxygenase genes detected in coal-tar-contaminated site undergoing active bioremediation. J. Appl. Microbiol . 108, 1252–1262. doi: 10.1111/j.1365-2672.2009.04523.x
Kumari, P., Meena, M., and Upadhyay, R. S. (2018). Characterization of plant growth promoting rhizobacteria (PGPR) isolated from the rhizosphere of Vigna radiata (mung bean). Biocatal. Agric. Biotechnol. 16, 155–162. doi: 10.1016/j.bcab.2018.07.029
Labie, D. (2007). Developmental neurotoxicity of industrial chemicals. Med. Sci. 23, 868–872. doi: 10.1051/medsci/20072310868
Lau, H. T., Suh, H. W., Golkowski, M., and Ong, S. E. (2014). Comparing SILAC- and stable isotope dimethyl-labeling approaches for quantitative proteomics. J. Proteome Res. 13, 4164–4174. doi: 10.1021/pr500630a
Lee, W. N., and Go, V. L. (2005). Nutrient-gene interaction: tracer-based metabolomics. J. Nutr. 135, 3027S−3032S. doi: 10.1093/jn/135.12.3027S
Liang, L., Stone, R. C., Stojadinovic, O., Ramirez, H., Pastar, I., Maione, A. G., et al. (2016). Integrative analysis of miRNA and mRNA paired expression profiling of primary fibroblast derived from diabetic foot ulcers reveals multiple impaired cellular functions. Wound Repair Regen . 24, 943–953. doi: 10.1111/wrr.12470
Lin, C. Y., Viant, M. R., and Tjeerdema, R. S. (2006). Metabolomics: methodologies and applications in the environmental sciences. J. Pestic. Sci. 31, 245–251. doi: 10.1584/jpestics.31.245
Liu, L., Li, Y. H., Li, S., Hu, N., He, Y., Pong, R., et al. (2012). Comparison of next-generation sequencing systems. J. Biomed. Biotechnol. 2012:251364. doi: 10.1155/2012/251364
Loman, N. J., and Pallen, M. J. (2015). Twenty years of bacterial genome sequencing. Nat. Rev. Microbiol. 13, 1–9. doi: 10.1038/nrmicro3565
Lovley, D. R. (2003). Cleaning up with genomics: applying molecular biology to bioremediation. Nat. Rev. Microbiol . 1, 35–44. doi: 10.1038/nrmicro731
Lovley, D. R., Phillips, E. J. P., Gorby, Y. A., and Landa, E. R. (1991). Microbial reduction of uranium. Nature 350, 413–416. doi: 10.1038/350413a0
Lu, H., Giordano, F., and Ning, Z. (2016). Oxford nanopore MinION sequencing and genome assembly. Genom. Proteom. Bioinfo. 14, 265–279. doi: 10.1016/j.gpb.2016.05.004
Lu, H., Que, Y., Wu, X., Guan, T., and Guo, H. (2019). Metabolomics deciphered metabolic reprogramming required for biofilm formation. Sci. Rep. 9:13160. doi: 10.1038/s41598-019-49603-1
Luciene, M. C., Helen, C. R., Luciana, M. C., Priscila, A. R., de Sousa Danielle, F. O. M., and Nívia, M. M. C. (2015). “Bioremediation of polluted waters using microorganisms,” in Advances in Bioremediation of Wastewater and Polluted Soil , ed N. Shiomi (London: IntechOpen), 1–22. doi: 10.5772/60770
Malik, S., Beer, M., Megharaj, M., and Naidu, R. (2008). The use of molecular techniques to characterize the microbial communities in contaminated soil and water. Environ. Int. 34, 265–276. doi: 10.1016/j.envint.2007.09.001
Malla, M. A., Dubey, A., Yadav, S., Kumar, A., Hashem, A., and Abd Allah, E. F. (2018). Understanding and designing the strategies for the microbe-mediated remediation of environmental contaminants using omics approaches. Front. Microbiol. 9:1132. doi: 10.3389/fmicb.2018.01132
Mallick, H., Franzosa, E. A., Mclver, L. J., Banerjee, S., Sirota-Madi, A., Kostic, A. D., et al. (2019). Predictive metabolomic profiling of microbial communities using amplicon or metagenomic sequences. Nat. Commun. 10:3136. doi: 10.1038/s41467-019-10927-1
Maphosa, F., de Vos, W. M., and Smidt, H. (2010a). Exploiting the ecogenomics toolbox for environmental diagnostics of organohalide-respiring bacteria. Trends Biotechnol. 28, 308–316. doi: 10.1016/j.tibtech.2010.03.005
Maphosa, F., Smidt, H., de Vos, W. M., and Roling, W. H. F. M. (2010b). Microbial community and metabolite dynamics of an anoxic dechlorinating bioreactor. Stud. Environ. Sci . 44, 4884–4890. doi: 10.1021/es903721s
Mardis, E. R. (2008). Next-generation DNA sequencing methods. Annu. Rev. Genomics Hum. Genet. 9, 387–402. doi: 10.1146/annurev.genom.9.081307.164359
Maroli, A. S., Gaines, T. A., Foley, M. E., Duke, S. O., Doğramacı, M., Anderson, J. V., et al. (2018). Omics in weed science: A perspective from genomics, transcriptomics, and metabolomics approaches. Weed Sci. 66, 681–695. doi: 10.1017/wsc.2018.33
Martínez-Esteso, M. J., Casado-Vela, J., Sellés-Marchart, S., Pedreño, M. A., and Bru-Martínez, R. (2014). Differential plant proteome analysis by isobaric tags for relative and absolute quantitation (iTRAQ). Methods Mol. Biol. 1072, 155–169. doi: 10.1007/978-1-62703-631-3_12
Mattarozzi, M., Manfredi, M., Montanini, B., Gosetti, F., Sanangelantoni, A. M., Marengo, E., et al. (2017). A metaproteomic approach dissecting major bacterial functions in the rhizosphere of plants living in serpentine soil. Anal. Bioanal. Chem. 409, 2327–2339. doi: 10.1007/s00216-016-0175-8
Maxam, A. M., and Gilbert, W. A. (1977). A new method for sequencing DNA. Proc. Natl. Acad. Sci. U.S.A. 74, 560–564. doi: 10.1073/pnas.74.2.560
McMahon, M. A., Prakash, T. P., Cleveland, D. W., Bennett, C. F., and Rahdar, M. (2018). Chemically modified Cpf1-CRISPR RNAs mediate efficient genome editing in mammalian cells. Mol. Ther. 26, 1228–1240. doi: 10.1016/j.ymthe.2018.02.031
Medić, A., Stojanović, K., Izrael-Živković, L., Beškoski, V., Lončarević, B., Kazazić, S., et al. (2019). A comprehensive study of conditions of the biodegradation of a plastic additive 2,6-di-tert-butylphenol and proteomic changes in the degrader Pseudomonas aeruginosa san ai. RSC Adv . 9, 23696–23710. doi: 10.1039/C9RA04298A
Meena, M., Aamir, M., Vikas, K., Swapnil, P., and Upadhyay, R. S. (2018). Evaluation of morpho-physiological growth parameters of tomato in response to Cd induced toxicity and characterization of metal sensitive NRAMP3 transporter protein. Environ. Exp. Bot . 148, 144–167. doi: 10.1016/j.envexpbot.2018.01.007
Meena, M., Divyanshu, K., Kumar, S., Swapnil, P., Zehra, A., Shukla, V., et al. (2019). Regulation of L-proline biosynthesis, signal transduction, transport, accumulation and its vital role in plants during variable environmental conditions. Heliyon 5:e02951. doi: 10.1016/j.heliyon.2019.e02952
Meena, M., Dubey, M. K., Swapnil, P., Zehra, A., Singh, S., Kumari, P., et al. (2017a). “The rhizosphere microbial community and methods of its analysis,” in Advances in PGPR Research , eds H. B. Singh, B. K. Sarma, and C. Keswani (Boston, MA: CAB International), 275–295.
Meena, M., Gupta, S. K., Swapnil, P., Zehra, A., Dubey, M. K., and Upadhyay, R. S. (2017b). Alternaria toxins: potential virulence factors and genes related to pathogenesis. Front. Microbiol. 8:1451. doi: 10.3389/fmicb.2017.01451
Meena, M., Prasad, V., and Upadhyay, R. S. (2017c). Evaluation of biochemical changes in leaves of tomato infected with Alternaria alternata and its metabolites. Vegetos 30:2. doi: 10.5958/2229-4473.2017.00020.9
Meena, M., Prasad, V., Zehra, A., Gupta, V. K., and Upadhyay, R. S. (2015). Mannitol metabolism during pathogenic fungal–host interactions under stressed conditions. Front. Microbiol. 6, 1019–1026. doi: 10.3389/fmicb.2015.01019
Meena, M., and Samal, S. (2019). Alternaria host-specific (HSTs) toxins: an overview of chemical characterization, target sites, regulation and their toxic effects. Toxicol. Rep. 6, 745–758. doi: 10.1016/j.toxrep.2019.06.021
Meena, M., and Swapnil, P. (2019). Regulation of WRKY genes in plant defense with beneficial fungus Trichoderma : current perspectives and future prospects. Arch. Phytopathol. Plant Protect. 52, 1–17. doi: 10.1080/03235408.2019.1606490
Meena, M., Swapnil, P., and Upadhyay, R. S. (2017d). Isolation, characterization and toxicological potential of tenuazonic acid, alternariol and alternariol monomethyl ether produced by Alternaria species phytopathogenic on plants. Sci. Rep. 7:8777. doi: 10.1038/s41598-017-09138-9
Meena, M., Swapnil, P., Zehra, A., Aamir, M., Dubey, M. K., and Upadhyay, R. S. (2017e). “Beneficial microbes for disease suppression and plant growth promotion,” in Plant-Microbe Interactions in Agro-Ecological Perspectives , eds D. Singh, H. Singh, and R. Prabha (Singapore: Springer), 395–432. doi: 10.1007/978-981-10-6593-4_16
Meena, M., Zehra, A., Dubey, M. K., Aamir, M., Gupta, V. K., and Upadhyay, R. S. (2016). Comparative evaluation of biochemical changes in tomato ( Lycopersicon esculentum Mill.) infected by Alternaria alternata and its toxic metabolites (TeA, AOH, and AME). Front. Plant Sci. 7:1408. doi: 10.3389/fpls.2016.01408
Misra, B. B., Langefeld, C. D., Olivier, M., and Cox, L. A. (2018). Integrated omics: tools, advances, and future approaches. J. Mol. Endocrinol . 62, R21–R45. doi: 10.1530/JME-18-0055
Morgan, X. C., Tickle, T. L., Sokol, H., Gevers, D., Devaney, K. L., Ward, D. V., et al. (2012). Dysfunction of the intestinal microbiome in inflammatory bowel disease and treatment. Genome Biol. 13:R79. doi: 10.1186/gb-2012-13-9-r79
Morris, R. M., Nunn, B. L., Frazar, C., Goodlett, D. R., Ting, Y. S., and Rocap, G. (2010). Comparative metaproteomics reveals ocean-scale shifts in microbial nutrient utilization and energy transduction. ISME J. 4, 673–685. doi: 10.1038/ismej.2010.4
Muccee, F., and Ejaz, S. (2020). Whole genome shotgun sequencing of POPs degrading bacterial community dwelling tannery effluents and petrol contaminated soil. Microbiol. Res . 238:126504. doi: 10.1016/j.micres.2020.126504
Muffler, A., Bettermann, S., Haushalter, M., Hörlein, A., Neveling, U., Schramm, M., et al. (2002). Genomewide transcription profiling of Corynebacterium glutamicum after heat shock and during growth on acetate and glucose. J. Biotechnol . 98, 255–268. doi: 10.1016/S0168-1656(02)00136-0
Nagalakshmi, U., Waern, K., and Snyder, M. (2010). RNA-Seq: a method for comprehensive transcriptome analysis. Curr. Protoc. Mol. Biol. 4, 1–13. doi: 10.1002/0471142727.mb0411s89
Ngara, T. R., and Zhang, H. (2018). Recent advances in function-based metagenomic screening. Genom. Proteom. Bioinf. 16, 405–415. doi: 10.1016/j.gpb.2018.01.002
Nikolaki, S., and Tsiamis, G. (2013). Microbial diversity in the era of omic technologies. Biomed. Res. Int. 2013, 958719. doi: 10.1155/2013/958719
Nojiri, H., and Tsuda, M. (2005). Functional evolution of bacteria in degradation of environmental pollutants. Tanpakushitsu Kakusan Koso 50, 1505–1509.
PubMed Abstract | Google Scholar
Oulas, A., Pavloudi, C., Polymenakou, P., Pavlopoulos, G. A., Papanikolaou, N., Kotoulas, G., et al. (2015). Metagenomics: tools and insights for analyzing next-generation sequencing data derived from biodiversity studies. Bioinform. Biol. Insights 9, 75–88. doi: 10.4137/BBI.S12462
Palmer, C., Bik, E. M., Eisen, M. B., Eckburg, P. B., Sana, T. R., Wolber, P. K., et al. (2006). Rapid quantitative profiling of complex microbial populations. Nucleic Acids Res. 34:e5. doi: 10.1093/nar/gnj007
Pan, Z., and Raftery, D. (2007). Comparing and combining NMR spectroscopy and mass spectrometry in metabolomics. Anal. Bioanal. Chem. 387, 525–527. doi: 10.1007/s00216-006-0687-8
Panagiotou, G., Kouskoumvekaki, I., Jónsdóttir, S. O., and Olsson, L. (2007). Monitoring novel metabolic pathways using metabolomics and machine learning: induction of the phosphoketolase pathway in Aspergillus nidulans cultivations. Metabolomics 3, 503–516. doi: 10.1007/s11306-007-0061-7
Pandey, A., Tripathi, P. H., Tripathi, A. H., Pandey, S. C., and Gangola, S. (2019). “Omics technology to study bioremediation and respective enzymes,” in Smart Bioremediation Technologies: Microbial Enzymes , ed P. Bhatt (New Delhi: Academic Press), 23–43. doi: 10.1016/B978-0-12-818307-6.00002-0
Pantaleo, M. A., Ravegnini, G., Astolfi, A., Simeon, V., Nannini, M., Saponara, M., et al. (2016). Integrating miRNA and gene expression profiling analysis revealed regulatory networks in gastrointestinal stromal tumors. Epigenomics 8, 1347–1366. doi: 10.2217/epi-2016-0030
Paul, D., Pandey, G., Pandey, J., and Jain, R. K. (2005). Accessing microbial diversity for bioremediation and environmental restoration. Trends Biotechnol. 23, 135–142. doi: 10.1016/j.tibtech.2005.01.001
Peabody, M. A., van Rossum, T., Lo, R., and Brinkman, F. S. L. (2015). Evaluation of shotgun metagenomics sequence classification methods using in silico and in vitro simulated communities. BMC Bioinformatics 16:363. doi: 10.1186/s12859-015-0788-5
Plewniak, F., Crognale, S., Rossetti, S., and Bertin, P. N. (2018). A genomic outlook on bioremediation: the case of arsenic removal. Front. Microbiol. 9:820. doi: 10.3389/fmicb.2018.00820
Poirier, I., Hammann, P., Kuhn, L., and Bertrand, M. (2013). Strategies developed by the marine bacterium Pseudomonas fluorescens BA3SM1 to resist metals: a proteome analysis. Aquat. Toxicol. 128, 215–232. doi: 10.1016/j.aquatox.2012.12.006
Popovic, A., Hai, T., Tchigvintsev, A., Hajighasemi, M., Nocek, B., Khusnutdinova, A. N., et al. (2017). Activity screening of environmental metagenomic libraries reveals novel carboxylesterase families. Sci. Rep. 7:44103. doi: 10.1038/srep44103
Pritchard, C. C., Cheng, H. H., and Tewari, M. (2012). MicroRNA profiling: approaches and considerations. Nat. Rev. Genet. 13, 358–369. doi: 10.1038/nrg3198
Quince, C., Walker, A. W., Simpson, J. T., Loman, N. J., and Segata, N. (2017). Shotgun metagenomics, from sampling to analysis. Nat. Biotechnol. 35, 833–844. doi: 10.1038/nbt.3935
Raghunandan, K., Kumar, A., Kumar, S., Permaul, K., and Singh, S. (2018). Production of gellan gum, an exopolysaccharide, from biodiesel-derived waste glycerol by Sphingomonas spp. 3 Biotech. 8:71. doi: 10.1007/s13205-018-1096-3
Rani, A., Porwal, S., Sharma, R., Kapley, A., Purohit, H. J., and Kalia, V. C. (2008). Assessment of microbial diversity in effluent treatment plants by culture dependent and culture independent approaches. Bioresour. Technol. 99, 7098–7107. doi: 10.1016/j.biortech.2008.01.003
Ranjan, R., Rani, A., Metwally, A., McGee, H. S., and Perkins, D. L. (2016). Analysis of the microbiome: advantages of whole genome shotgun versus 16S amplicon sequencing. Biochem. Biophys. Res. Commun. 469, 967–977. doi: 10.1016/j.bbrc.2015.12.083
Rashid, M., and Stingl, U. (2015). Contemporary molecular tools in microbial ecology and their application to advancing biotechnology. Biotechnol. Adv. 33, 1755–1773. doi: 10.1016/j.biotechadv.2015.09.005
Reuter, J. A., Spacek, D. V., and Snyder, M. P. (2015). High-throughput sequencing technologies. Mol. Cell 58, 586–597. doi: 10.1016/j.molcel.2015.05.004
Rhee, S. K., Liu, X., Wu, L., Chong, S. C., Wan, X., and Zhou, J. (2004). Detection of genes involved in biodegradation and biotransformation in microbial communities by using 50-mer oligonucleotide microarrays. Appl. Environ. Microbiol. 70, 4303–4317. doi: 10.1128/AEM.70.7.4303-4317.2004
Rhoads, A., and Au, K. F. (2015). PacBio sequencing and its applications. Genom. Proteom. Bioinform. 13, 178–289. doi: 10.1016/j.gpb.2015.08.002
Riesenfeld, C. S., Schloss, P. D., and Handelsman, J. (2004). Metagenomics: genomic analysis of microbial communities. Ann. Rev. Genet. 38, 525–552. doi: 10.1146/annurev.genet.38.072902.091216
Rodríguez, A., Castrejón-Godínez, M. L., Salazar-Bustamante, E., Gama-Martínez, Y., Sánchez-Salinas, E., Mussali-Galante, P., et al. (2020). Omics approaches to pesticide biodegradation. Curr. Microbiol. 77, 545–563. doi: 10.1007/s00284-020-01916-5
Rogers, S. L., and McClure, N. (2003). “The role of microbiological studies in bioremediation process optimization,” in Bioremediation: A Critical Review , eds I. M. Head, I. Singleton, and M. Milner (Norfolk: Horizon Scientific Press), 27–59.
Ronaghi, M., Karamohamed, S., Pettersson, B., Uhlén, M., and Nyrén, P. (1996). Real-time DNA sequencing using detection of pyrophosphate release. Anal. Biochem. 242, 84–89. doi: 10.1006/abio.1996.0432
Roume, H., Heintz-Buschart, A., Muller, E. E. L., May, P., Satagopam, V. P., Laczny, C. C., et al. (2015). Comparative integrated omics: identification of key functionalities in microbial community-wide metabolic networks. NPJ Biofilms Microb . 1:15007. doi: 10.1038/npjbiofilms.2015.7
Sakshi and Haritash A. K. (2020). A comprehensive review of metabolic and genomic aspects of PAH-degradation. Arch. Microbiol. 202, 2033–2058. doi: 10.1007/s00203-020-01929-5
Samorodnitsky, E., Jewell, B. M., Hagopian, R., Miya, J., Wing, M. R., Lyon, E., et al. (2015). Evaluation of hybridization capture versus amplicon-based methods for whole-exome sequencing. Hum. Mutat . 36, 903–914. doi: 10.1002/humu.22825
Sanger, F., and Coulson, A. R. (1975). A rapid method for determining sequences in DNA by primed synthesis with DNA polymerase. J. Mol. Biol. 94, 441–448. doi: 10.1016/0022-2836(75)90213-2
Sato, Y., Hori, T., Koike, H., Navarro, R. R., Ogata, A., and Habe, H. (2019). Transcriptome analysis of activated sludge microbiomes reveals an unexpected role of minority nitrifiers in carbon metabolism. Commun. Biol . 2:179. doi: 10.1038/s42003-019-0418-2
Sauer, U. (2006). Metabolic networks in motion: 13C-based flux analysis. Mol. Syst. Biol. 2:62. doi: 10.1038/msb4100109
Schneegurt, M. A., and Kulpa, C. F. (1998). The application of molecular techniques in environmental biotechnology for monitoring microbial systems. Biotechnol. Appl. Biochem. 27, 73–79. doi: 10.1111/j.1470-8744.1998.tb01377.x
Scholz, M. B., Lo, C. C., and Chain, P. S. (2012). Next generation sequencing and bioinformatic bottlenecks: the current state of metagenomic data analysis. Curr. Opin. Biotechnol. 23, 9–15. doi: 10.1016/j.copbio.2011.11.013
Schut, G. J., Zhou, J., and Adams, M. W. (2001). DNA microarray analysis of the hyperthermophilic archaeon Pyrococcus furiosus : evidence for a new type of sulfur-reducing enzyme complex. J. Bacteriol. 183, 7027–7036. doi: 10.1128/JB.183.24.7027-7036.2001
Seo, J. S., Keum, Y. S., and Li, Q. X. (2009). Bacterial degradation of aromatic compounds. Int. J. Environ. Res. Public Health 6, 278–309. doi: 10.3390/ijerph6010278
Shah, V., Zakrzewski, M., Wibberg, D., Eikmeyer, F., Schlüter, A., and Madamwar, D. (2013). Taxonomic profiling and metagenome analysis of a microbial community from a habitat contaminated with industrial discharges. Microb. Ecol. 66, 533–550. doi: 10.1007/s00248-013-0253-9
Sharmin, F., Wakelin, S., Huygens, F., and Hargreaves, M. (2013). Firmicutes dominate the bacterial taxa within sugar-cane processing plants. Sci. Rep. 3:3107. doi: 10.1038/srep03107
Sharpton, T. J. (2014). An introduction to the analysis of shotgun metagenomic data. Front. Plant Sci . 5:209. doi: 10.3389/fpls.2014.00209
Shendure, J. (2008). The beginning of the end for microarrays? Nat. Methods 5, 585–587. doi: 10.1038/nmeth0708-585
Shendure, J., Porreca, G. J., Reppas, N. B., Lin, X., McCutcheon, J. P., and Rosenbaum, A. M. (2005). Accurate multiplex polony sequencing of an evolved bacterial genome. Science 309, 1728–1732. doi: 10.1126/science.1117389
Shi, Z. J., Yin, H., Van Nostrand, J. D., Voordeckers, J. W., Tu, Q., Deng, Y., et al. (2019). Functional gene array-based ultrasensitive and quantitative detection of microbial populations in complex communities. mSystems 4:e00296–e00219. doi: 10.1128/mSystems.00296-19
Siggins, A., Gunnigle, E., and Abram, F. (2012). Exploring mixed microbial community functioning: recent advances in metaproteomics. FEMS Microbiol. Ecol. 80, 265–280. doi: 10.1111/j.1574-6941.2011.01284.x
Silva, C. C., Hayden, H., Sawbridge, T., Mele, P., De Paula, S. O., Silva, L. C. F., et al. (2013). Identification of genes and pathways related to phenol degradation in metagenomic libraries from petroleum refinery wastewater. PLoS ONE 8:e61811. doi: 10.1371/journal.pone.0061811
Simmons, S. L., Dibartolo, G., Denef, V. J., Aliaga Goltsman, D. S., Thelen, M. P., and Banfield, J. F. (2008). Population genomic analysis of strain variation in Leptospirillum group II bacteria involved in acid mine drainage formation. PLoS Biol. 6:e177. doi: 10.1371/journal.pbio.0060177
Singh, J., Behal, A., Singla, N., Joshi, A., Birbian, N., Singh, S., et al. (2009). Metagenomics: Concept, methodology, ecological inference and recent advances. Biotechnol. J. 4, 480–494. doi: 10.1002/biot.200800201
Sueoka, K., Satoh, H., Onuki, M., and Mino, T. (2009). Microorganisms involved in anaerobic phenol degradation in the treatment of synthetic coke-oven wastewater detected by RNA stable-isotope probing. FEMS Microbiol. Lett. 291, 169–174. doi: 10.1111/j.1574-6968.2008.01448.x
Tang, Y. J., Hwang, J. S., Wemmer, D. E., and Keasling, J. D. (2007). Shewanella oneidensis MR-1 fluxome under various oxygen conditions. Appl. Environ. Microbiol. 73, 718–729. doi: 10.1128/AEM.01532-06
Thakur, M., Medintz, I. L., and Walper, S. A. (2019). Enzymatic bioremediation of organophosphate compounds—progress and remaining challenges. Front. Bioeng. Biotechnol. 7, 289. doi: 10.3389/fbioe.2019.00289
Turcatti, G., Romieu, A., Fedurco, M., and Tairi, A. P. (2008). A new class of cleavable fluorescent nucleotides: synthesis and optimization as reversible terminators for DNA sequencing by synthesis. Nucleic Acids Res. 36:e25. doi: 10.1093/nar/gkn021
van Nostrand, J. D., He, Z., and Zhou, J. (2011). “Dynamics of microbes in the natural setting: development of the Geochip,” in Environmental Microbiology: Current Technology and Water Applications , eds K. Sen and N. J. Ashbolt (Norfolk: Caister Academic Press), p. 316.
van Rossum, T., Kengen, S. W., and van der Oost, J. (2013). Reporter-based screening and selection of enzymes. FEBS J. 280, 2979–2996. doi: 10.1111/febs.12281
Vandera, E., Samiotaki, M., Parapouli, M., Panayotou, G., and Koukkou, A. I. (2015). Comparative proteomic analysis of Arthrobacter phenanthrenivorans Sphe3 on phenanthrene, phthalate and glucose. J. Proteom. 113, 73–89. doi: 10.1016/j.jprot.2014.08.018
Venter, J. C., Remington, K., Heidelberg, J. F., Halperna, A. L., Rusch, D., Eisen, J. A., et al. (2004). Environmental genome shotgun sequencing of the Sargasso Sea. Science 304, 66–74. doi: 10.1126/science.1093857
Verberkmoes, N. C., Denef, V. J., Hettich, R. L., and Banfield, J. F. (2009). Systems biology: functional analysis of natural microbial consortia using community proteomics. Nat. Rev. Microbiol. 7, 196–205. doi: 10.1038/nrmicro2080
Vermote, L., Verce, M., de Vuyst, L., and Weckx, S. (2018). Amplicon and shotgun metagenomic sequencing indicates that microbial ecosystems present in cheese brines reflect environmental inoculation during the cheese production process. Int. Dairy J. 87, 44–53. doi: 10.1016/j.idairyj.2018.07.010
Viant, M. R., and Sommer, U. (2013). Mass spectrometry based environmental metabolomics: a primer and review. Metabolomics 9, 144–158. doi: 10.1007/s11306-012-0412-x
Vigneron, A., Alsop, E. B., Cruaud, P., Philibert, G., King, B., Baksmaty, L., et al. (2017). Comparative metagenomics of hydrocarbon and methane seeps of the Gulf of Mexico. Sci. Rep. 7:16015. doi: 10.1038/s41598-017-16375-5
Villas-Bôas, S. G., Nielsen, J., Smedsgaard, J., Hansen, M. A. E., and Roessner-Tunali, U. (2007). Metabolome Analysis: An Introduction . Hoboken, NJ: Wiley-Interscience.
Vos, M., and Velicer, G. J. (2006). Genetic population structure of the soil bacterium Myxococcus xanthus at the centimeter scale. Appl. Environ. Microbiol. 72, 3615–3625. doi: 10.1128/AEM.72.5.3615-3625.2006
Wang, D. Z., Kong, L. F., Li, Y. Y., and Xie, Z. X. (2016). Environmental microbial community proteomics: status, challenges and perspectives. Int. J. Mol. Sci. 17, 1275. doi: 10.3390/ijms17081275
Wang, D. Z., Xie, Z. X., and Zhang, S. F. (2014). Marine metaproteomics: Current status and future directions. J. Proteom. 97, 27–35. doi: 10.1016/j.jprot.2013.08.024
Wang, J. H., Byun, J., and Pennathur, S. (2010). Analytical approaches to metabolomics and applications to systems biology. Semin. Nephrol. 30, 500–511. doi: 10.1016/j.semnephrol.2010.07.007
Ward, M. A., Hyman, B., Scherzer, C., Trisini-Lipsanopoulos, A., Ivinson, A. J., Lößner, C., et al. (2014). Exploration of plasma biomarkers for alzheimer's disease using isotopic tandem mass tags and a combined directed/data-dependent acquisition NLC-MS/MS method. Alzheimers Dement. J. Alzheimers Assoc. 10:P674. doi: 10.1016/j.jalz.2014.05.1215
Wiechert, W. (2001). 13C metabolic flux analysis. Metab. Eng. 3, 195–206. doi: 10.1006/mben.2001.0187
Wiechert, W., Möllney, M., Petersen, S., and de Graaf, A. A. (2001). A universal framework for 13C metabolic flux analysis. Metab. Eng. 3, 265–283. doi: 10.1006/mben.2001.0188
Wierckx, N. J. P., Ballerstedt, H., De Bont, J. A. M., DeWinde, J. H., Ruijssenaars, H. J., and Wery, J. (2008). Transcriptome analysis of a phenol-producing Pseudomonas putida S12 construct: genetic and physiological basis for improved production. J. Bacteriol. 190, 2822–2830. doi: 10.1128/JB.01379-07
Wierckx, N. J. P., Ruijssenaars, H. J., deWinde, J. H., Schmid, A., and Blank, L. M. (2009). Metabolic flux analysis of a phenol producing mutant of Pseudomonas putida S12: verification and complementation of hypotheses derived from transcriptomics. J. Biotechnol. 143, 124–129. doi: 10.1016/j.jbiotec.2009.06.023
Wilmes, P., and Bond, P. L. (2006). Metaproteomics: studying functional gene expression in microbial ecosystems. Trends Microbiol. 14, 92–97. doi: 10.1016/j.tim.2005.12.006
Wiseschart, A., Mhuantong, W., Tangphatsornruang, S., Chantasingh, D., and Pootanakit, K. (2019). Shotgun metagenomic sequencing from Manao-Pee cave, Thailand, reveals insight into the microbial community structure and its metabolic potential. BMC Microbiol. 19:144. doi: 10.1186/s12866-019-1521-8
Wong, D. W. S. (2018). Gene Targeting and Genome Editing. The ABCs of Gene Cloning . Cham: Springer, 187–197. doi: 10.1007/978-3-319-77982-9_20
Wright, R. J., Bosch, R., Gibson, M. I., and Christie-Oleza, J. A. (2020). Plasticizer degradation by marine bacterial isolates: a proteogenomic and metabolomic characterization. Environ. Sci. Technol. 54, 2244–2256. doi: 10.1021/acs.est.9b05228
Wrighton, K. C., Thomas, B. C., Sharon, I., Miller, C. S., Castelle, C. J., and VerBerkmoes, N. C. (2012). Fermentation, hydrogen, and sulfur metabolism in multiple uncultivated bacterial phyla. Science 337, 1661–1665. doi: 10.1126/science.1224041
Wu, C. C., and Yates, J. R. (2003). The application of mass spectrometry to membrane proteomics. Nat. Biotechnol. 21, 262–267. doi: 10.1038/nbt0303-262
Wu, Y. R., Luo, Z. H., Kwok-Kei Chow, R., and Vrijmoed, L. L. P. (2010). Purification and characterization of an extracellular laccase from the anthracene-degrading fungus Fusarium solani MAS2. Bioresour. Technol. 101, 9772–9777. doi: 10.1016/j.biortech.2010.07.091
Xie, J., He, Z., Liu, X., Liu, X., Van Nostrand, J. D., Deng, Y., et al. (2011). GeoChip-based analysis of the functional gene diversity and metabolic potential of microbial communities in acid mine drainage. Appl. Environ. Microbiol . 77, 991–999. doi: 10.1128/AEM.01798-10
Xue, K., Yuan, M. M., Shi, Z. J., Qin, Y., Deng, Y., Cheng, L., et al. (2016). Tundra soil carbon is vulnerable to rapid microbial decomposition under climate warming. Nat. Clim. Chang. 6:595. doi: 10.1038/nclimate2940
Yao, C., Chen, B. H., Joehanes, R., Otlu, B., Zhang, X., Liu, C., et al. (2015). Integromic analysis of genetic variation and gene expression identifies networks for cardiovascular disease phenotypes. Circulation 131, 536–549. doi: 10.1161/CIRCULATIONAHA.114.010696
Yergeau, E., Michel, C., Tremblay, J., Niemi, A., King, T. L., Wyglinski, J., et al. (2017). Metagenomic survey of the taxonomic and functional microbial communities of seawater and sea ice from the Canadian Arctic. Sci. Rep. 7:42242. doi: 10.1038/srep42242
Yoneda, A., Henson, W. R., Goldner, N. K., Park, K. J., Forsberg, K. J., Kim, S. J., et al. (2016). Comparative transcriptomics elucidates adaptive phenol tolerance and utilization in lipid-accumulating Rhodococcus opacus PD630. Nucleic Acids Res. 44, 2240–2254. doi: 10.1093/nar/gkw055
Yozwiak, N. L., Skewes-Cox, P., Stenglein, M. D., Balmaseda, A., Harris, E., and DeRisi, J. L. (2012). Virus identification in unknown tropical febrile illness cases using deep sequencing. PLoS Negl. Trop. Dis. 6:e1485. doi: 10.1371/journal.pntd.0001485
Zehra, A., Meena, M., Swapnil, P., Raytekar, N. A., and Upadhyay, R. S. (2020). “Sustainable approaches to remove heavy metals from water,” in Microbial Biotechnology: Basic Research and Applications , eds J. Singh, A. Vyas, S. Wang, and R. Prasad (Singapore: Springer), 127–146. doi: 10.1007/978-981-15-2817-0_6
Zhang, J., Chiodini, R., Badr, A., and Zhang, G. (2011). The impact of next-generation sequencing on genomics. J. Genet. Genom. 38, 95–109. doi: 10.1016/j.jgg.2011.02.003
Zhang, W., Li, F., and Nie, L. (2010). Integrating multiple 'omics' analysis for microbial biology: application and methodologies. Microbiology 156, 287–301. doi: 10.1099/mic.0.034793-0
Zhong, M., and Zhou, Q. (2002). Molecular-ecological technology of microorganisms and its application to research on environmental pollution. Ying Yong Sheng Tai Xue Bao 13, 247–251.
Zhou, J., He, Z., Yang, Y., Deng, Y., Tringe, S. G., and Alvarez-Cohen, L. (2015). High-throughput metagenomic technologies for complex microbial community analysis: open and closed formats. mBio 6:e02288-14. doi: 10.1128/mBio.02288-14
Zhou, J., Kang, S., Schadt, C. W., and Garten, C. T. (2008). Spatial scaling of functional gene diversity across various microbial taxa. Proc. Natl. Acad. Sci. U.S.A. 105, 7768–7773. doi: 10.1073/pnas.0709016105
Zhuang, X., Han, Z., Bai, Z., Zhuang, G., and Shim, H. (2010). Progress in decontamination by halophilic microorganisms in saline wastewater and soil. Environ. Pollut. 158, 1119–1126. doi: 10.1016/j.envpol.2010.01.007
Keywords: bioremediation, microorganisms, microarrays, metagenomics, transcriptomics, proteomics, metabolomics, fluxomics
Citation: Chandran H, Meena M and Sharma K (2020) Microbial Biodiversity and Bioremediation Assessment Through Omics Approaches. Front. Environ. Chem. 1:570326. doi: 10.3389/fenvc.2020.570326
Received: 07 June 2020; Accepted: 18 August 2020; Published: 25 September 2020.
Reviewed by:
Copyright © 2020 Chandran, Meena and Sharma. This is an open-access article distributed under the terms of the Creative Commons Attribution License (CC BY) . The use, distribution or reproduction in other forums is permitted, provided the original author(s) and the copyright owner(s) are credited and that the original publication in this journal is cited, in accordance with accepted academic practice. No use, distribution or reproduction is permitted which does not comply with these terms.
*Correspondence: Mukesh Meena, mukeshmeenamlsu@gmail.com ; mukeshmeenabhu@gmail.com
- Review Article
- Published: 21 September 2012
Application of bioremediation technology in the environment contaminated with petroleum hydrocarbon
- Subhash Chandra 1 ,
- Richa Sharma 1 ,
- Kriti Singh 1 &
- Anima Sharma 2
Annals of Microbiology volume 63 , pages 417–431 ( 2013 ) Cite this article
17k Accesses
129 Citations
Metrics details
The demand for petroleum and petroleum products is increasing day by day, while oil spills and improper discharge of industrial wastes contributes to the rising contamination of the environment with petroleum hydrocarbon. Petroleum hydrocarbon spills cause various carcinogenic and neurotoxic effects, and thus effective treatment strategies are required. The various physical and chemical methods currently in use are costly and leave toxic residues in the environment. In contrast, bioremediation is a promising technology in the treatment of petroleum hydrocarbon contamination because of its high effectiveness, lower cost and environmental synergy. Here, we review the biodegradation of petroleum hydrocarbon and various factors influencing this process.
Introduction
As the world’s population continues to grow, the demand for petroleum and petroleum products increases vigorously, while oil spills and the improper discharge of industrial wastes contribute to pollution of the environment (Hernandez et al. 1998 ; Gupta and Mahapatra 2003 ; Strong and Burgess 2008 ). In 1971, in Whitemarsh Township, Pennsylvania, a gasoline pipeline broke and released several hundred thousand gallons of high-octane gasoline. It was estimated that 100,000 gallons (379,000 L) of gasoline seeped into the underlying dolomitic aquifer and the dissolved petroleum hydrocarbons migrated to a nearby municipal water supply well. It was recently proposed that at the time, that ground water system might have contained petroleum hydrocarbon-degrading microorganisms (Litchfield and Clark 1973 ). Another majoroil spill occured in 1989, in Prince William Sound, Alaska, in which the oil tanker Exxon Valdez wrecked and spilled more than 200,000 barrels of crude oil into the water (Hagar 1989 ). Several other major oil spills have occurred in Texas, Rhode Island, and the Delaware Bay (Anonymous 1989 ). These spills have attracted the attention of researchers examining the problem of hydrocarbon contamination and remediation of water and land environments. The amount of natural crude oil seepage was estimated to be 600,000 tons per year, with a range of uncertainty of 200,000 metric tons per year (Kvenvolden and Cooper 2003 ). Thus, control and treatment strategies to reduce the hazardous effects of oil pollution are needed. However, conventional physical-chemical treatments have high costs and can generate residues that are toxic to the biota. Applying high efficiency and low cost bioremediation processes represent an extremely important way of recovering contaminated areas among several other clean up techniques (Bidoia et al. 2010 ). The success of bioremediation efforts in the cleanup of the Exxon Valdez oil spill created tremendous interest in the potential of biodegradation and bioremediation technology. This was one of the first instances where bioremediation was actively considered as a remedial strategy to clean up a petroleum-contaminated ground system (Chapelle 1999 ). Bioremediation, a process that utilizes the natural capability of microorganisms to degrade toxic waste, is emerging as a promising technology for the treatment of soil and groundwater contamination (Milic et al. 2009 ). This review contains a broad and updated overview on bioremediation of hydrocarbon contaminated-environments, microbial degradation of petroleum hydrocarbons, and various physical, chemical, and biological factors influencing biodegradation.
- Petroleum hydrocarbon
Petroleum was first obtained by the Chinese in pre-Christian times and has been known for several years to occur in surface seepage (Okoh 2006 ). The modern petroleum industry had its beginning in Romania and in a Pennsylvania well sunk by Colonel E. A. Drake in 1859 (Alloway and Ayres 1993 ). Petroleum is a naturally occurring, oily, flammable liquid that is present beneath the surface of the earth. It is a mixture of hydrocarbons and elements like sulphur, oxygen, and nitrogen. In the refining industry, petroleum refers only to crude oil (Van 1997 ). Crude oil originates in naturally occurring geological deposits formed from the organic decomposition of ancient plants and animals under high temperatures and pressure. Crude oil is principally composed of liquid hydrocarbons (HCs) having four or more carbon atoms (Manning and Thompson 1991 ; Van 1997 ). Alkanes, cycloalkanes,and aromatics are the three principle classes of hydrocarbons present in petroleum (Strausz and Lown 2003 ). Alkenes and alkynes are present only in trace quantities (Van 1997 ; Potter and Simmons 1998 ). These compounds can be categorized into four simple fractions: saturated (or alkanes); aromatics, including compounds such as benzene, toluene, ethylbenzene and xylenes; (BTEX) and polyaromatic hydrocarbons (PAHs); resins, consisting of compounds containing nitrogen, sulphur, and oxygen that are dissolved in oil; and asphaltenes, which are large and complex molecules that are colloidally dispersed in oil. The physical properties and exact chemical composition of crude oil varies from one locality to another. In most soil and sediments, hydrocarbons are present in low concentrations (Rosenberg et al. 1992 ). Hydrocarbon contamination of surface and subsurface soil can result from accidental spills, equipment failure, and leaks in underground facilities, such as underground storage tanks, pipeline, or earthen pits (flare pits, sumps). At abandoned well sites, the source of PHCs in soils is generally from spills at well centers or from the disposal of drilling wastes in sumps or flare pits that are used to store or burn fluids produced at the sites. Soil contamination with hydrocarbons causes extensive damage of the local system since bioaccumulation of pollutants in animals and plant tissue may cause death or mutations (Alvarez and Vogel 1991 ).
Hazards of petroleum hydrocarbon contamination
The primary effects of exposure to petroleum hydrocarbons are central nervous system depression and polyneuropathy. Studies have shown that repeated exposure of laboratory animals to high concentrations of whole gasoline vapors at 67,262 and 2,056 ppm caused kidney damage and kidney cancer in rats and liver cancer in mice. The International Agency for Research on Cancer (IARC) has listed gasoline as possibly carcinogenic (2B) to humans (IARC 2000 ; Mortelmans et al. 1986 ). Among the components of gasoline, N-heptane and cyclohexane are known to cause narcosis and irritation of the eyes and mucous membranes. Cyclohexane has been reported to cause liver and kidney changes in rabbits, and n-heptane has been reported to cause polyneuritis following prolonged exposure. The IARC, the National Toxicology program (NTP) and the Occupational Safety and Health Adminstration (OSHA) all classify benzene as a human carcinogen. Studies have shown that the effects of benzene exposure can include leukemia at concentrations as low as 1 ppm. Methyl tertiary butyl ester (MTBE) is another potential human carcinogen, with studies showing that this component is a carcinogen in animals. It does not appear to readily biodegrade in the environment and persists both in aerobic and anaerobic conditions; it has an unpleasant taste and odor even at low concentrations. Other compounds of interest in petroleum are benzene toluene, ethyl benzene, and xylenes (BTEX). The isomers of trimethylbenzene may cause nervousness, tension, anxiety and asthmatic bronchitis. n-Hexane is known to cause the polyneuropathy (peripheral nerve damage) after repeated and prolonged exposure, while other hexanes show narcotic effects at 1,000 ppm and are not metabolized like n-hexane. Toluene can cause impairment of coordination and momentary loss of memory at exposures of 200–500 ppm, and causes palpations at 500–1,500 ppm. Crude oil is not considered a hazardous waste (petroleum is considered a hazardous material), but crude oil coats and kills sea life and mucks up the surrounding beaches, rocks, trees, etc. (Viswanathan 1999 ).
Remediation processes
Physical and chemical processes are usually used in the remediation of contaminated areas; recovery, dispersion, dilution, sorption, volatilization and abiotic transformations are all important means of hydrocarbon elimination. However, these types of treatment systems require heavy machinery and the environmental consequences of removing pollutants with these techniques may result in massive air pollution (Matsumiya and Kubo 2007 ). Allying high efficiency and low costs, bioremediation processes represent an extremely important alternative for rehabilitating oil-contaminated areas compared to other conventional clean up techniques (Bhupathiraju et al. 2002 ). Remediation by physical treatments has shown that even though the soil is unaffected or unmodified by these treatments, the costs are high and thus not applicable on a large scale. In spite of this, biodegradation is most often the primary mechanism for contaminant clean-up (Alexander 1999 ; Margesin and Schinner 2001 ) and for the remediation of hydrocarbon-contaminated sites, biological technologies are a widely used, cost-efficient and sustainable approach (Leeson and Hinchee 1997 ).
- Bioremediation
Bioremediation, which utilizes the biodegradative potentials of organisms or their attributes, is an effective technology that can be used to accomplish both effective detoxification and volume reduction. It is useful in the recovery of sites contaminated with oil and hazardous wastes (Caplan 1993 ). Moreover, bioremediation technology is believed to be noninvasive and relatively cost-effective (April et al. 2000 ). In some cases, it may not require more than the addition of some degradation enhancers (surfactants, nutrients, etc.) to the polluted system. It could end up being the most reliable and probably least expensive option for exploitation in solving some chemical pollution problems (Mesarch et al. 2000 ).

Physical factors affecting biodegradation of petroleum hydrocarbon
Knowledge of the characters and parameters that affect the microbial biodegradation of pollutants is required for the successful application of bioremediation technology. A number of limiting factors have been recognized that affect the biodegradation of petroleum hydrocarbons, many of which have been discussed by Brusseau ( 1998 ).
Physical and chemical composition of hydrocarbon
The susceptibility of hydrocarbons to biodegradation is determined by the structure and molecular weight of the hydrocarbon molecule. Aliphatic hydrocarbons are degraded and assimilated by a wide range of microorganisms. n-alkanes of intermediate chain length (C 10 –C 24 ) are degraded most rapidly. Short-chain alkanes (less than C9) are toxic to many microorganisms but being volatile, are generally lost rapidly in the atmosphere. Long-chain alkanes are generally resistant to biodegradation. Branching in general reduces the rate of biodegradation. Aromatics may be partly oxidized but are metabolized by only a few bacteria. Aromatic compounds, especially polyaromatic hydrocarbons (PAHs), are degraded slowly. Alicyclic compounds can be degraded via a process known as cometabolism. The bioavailability of hydrocarbons, which is largely a function of concentration and physical state, as well as their hydrophobicity, sorption onto soil particles, volatilization and solubility greatly affect the extent of their biodegradation. Hydrocarbons differ in their susceptibility to microbial attack and, in the past, have generally been ranked in the following order of decreasing susceptibility: n-alkanes >branched alkanes >low-molecular-weight aromatics >cyclic alkanes (Perry 1984 ). Biodegradation rates have been shown to be highest for the saturated, followed by the light aromatics, with high-molecular-weight aromatics and polar compounds exhibiting extremely low rates of degradation (Jobson et al. 1972 ; Walker et al. 1976 ; Fusey and Oudot 1984 ). This pattern is not universal, however, as Cooney et al. ( 1985 ) reported observing greater degradation of naphthalene instead of hexadecane in water-sediment mixtures from a freshwater lake and extensive biodegradation of alkylaromatics in marine sediments prior to detectable changes in the n-alkane profile of crude oil (Jones et al. 1983 ). Fedorak and Westlake ( 1981 ) also reported a more rapid attack of aromatic hydrocarbons during the degradation of crude oil by marine microbial populations from a pristine site and a commercial harbor.
Temperature
Among physical factors, temperature plays an important role in the biodegradation of hydrocarbons by directly affecting the chemistry of the pollutants as well as affecting the physiology and diversity of the microbial flora. Atlas ( 1975 ) found that at low temperatures, the viscosity of the oil increased, while the volatility of the toxic low molecular weight hydrocarbons were reduced, delaying the onset of biodegradation. Temperature also affects the solubility of hydrocarbons (Foght et al. 1996 ). At low temperature, hydrocarbon water solubility increases. Some previously reported data on the solubility of hydrocarbons in water are summarized in Table 1 (Bahram 1969 ) and 2 ( http://esc.syrres.com/efdb/chemfate.htm ). Although hydrocarbon biodegradation can occur over a wide range of temperatures, rates of degradation are generally observed to decrease with decreasing temperature; this is believed to be a result primarily of decreased rates of enzymatic activity, or the “Q lo ” effect (Atlas and Bartha 1972 ; Gibbs et al. 1975 ). The highest degradation rates generally occur in the range of 30–40°C in soil environments, 20–30°C in some freshwater environments and 15–20°C in marine environments (Bartha and Bossert 1984 ; Cooney 1984 ). Venosa and Zhu ( 2003 ) reported that ambient temperatures of the environment affected both the properties of spilled oil and the activity of the microorganisms. Significant biodegradation of hydrocarbons have been reported in psychrophilic environments in temperate regions (Pelletier et al. 2004 ; Delille et al. 2004 ), however, the bulk of information that exists on hydrocarbon degradation details the activities of mesophiles (Yumoto et al. 2002 ; Pelletier et al. 2004 ; Delille et al. 2004 ). Documented research on the environmental consequences of terrestrial spills in cold regions remains scarce, even though petroleum contamination is recognized as a significant threat to polar environments. Full-scale in situ remediation of petroleum contaminated soils has not yet been used in Antarctica for example, partly because it has long been assumed that air and soil temperatures are too low for effective biodegradation (Delille et al. 2004 ). Climate and season would be expected to select for different populations of hydrocarbon-utilizing microorganisms which are adapted to ambient temperatures. Colwell et al. ( 1978 ) reported extensive degradation of Metula crude oil by mixed cultures of marine bacteria at 3 °C, and Huddleston and Cresswell ( 1976 ) observed petroleum biodegradation in soil at −1.1 °C. By contrast, only negligible degradation of oil was exhibited in frozen tundra soil (Atlas et al. 1976 ) and in Arctic marine ice (Atlas et al. 1978 ). Low winter temperatures were the limiting factor for the biodegradation of polyaromatic hydrocarbons in estuarine sediment (Shiaris 1989 ) and of a variety of hydrocarbons in freshwater lakes (Cooney et al. 1985 ).
Hydrocarbons, being highly reduced substrates, require an electron acceptor, with molecular oxygen being most common. Though most studies have shown the biodegradation of hydrocarbons to be an aerobic process, anaerobic biodegradation of hydrocarbons has also been reported. In the absence of molecular oxygen, nitrate, iron, bicarbonate, nitrous oxide and sulfate have been shown to act as an alternate electron acceptors during hydrocarbon degradation. Oxygen is typically the limiting factor in aerobic bioremediation at many sites. The degradation of petroleum hydrocarbons occurs much faster under aerobic conditions compared to anaerobic conditions. Therefore, the addition of oxygen can significantly increase the remediation rate. The initial steps in the catabolism of aliphatic (Singer and Finnerty 1984a ), cyclic (Perry 1984 ), and aromatic (Cerniglia 1984a ) hydrocarbons by bacteria and fungi involve the oxidation of the substrate by oxygenases, for which molecular oxygen is required. The concentration of oxygen has been identified as the rate-limiting variable in the biodegradation of petroleum in soil (Von Wedel et al. 1988 ) and of gasoline in groundwater (Jamison et al. 1975 ). The availability of oxygen in soils is dependent upon rates of microbial oxygen consumption, the type of soil, whether the soil is waterlogged, and the presence of utilizable substrates which can lead to oxygen depletion (Bossert and Bartha 1984 ). Anaerobic degradation of petroleum hydrocarbons by microorganisms has been shown in some studies to occur only at negligible rates (Bailey et al. 1973 ; Jamison et al. 1975 ; Ward and Brock 1978a ), and its ecological significance has been generally considered to be minor (Atlas 1981 ; Bossert and Bartha 1984 ; Cooney 1984 ; Floodgate 1984 ). However, the microbial degradation of oxidized aromatic compounds such as benzoate (Tarvin and Buswell 1934 ) and of halogenated aromatic compounds such as the halobenzoates (Suflita et al. 1982 ), chlorophenols (Boyd and Shelton 1984 ), and polychlorinated biphenyls (Chen et al. 1988 ) has been shown to occur under anaerobic conditions. Recent evidence also indicates that microbial consortia from soil and sludge are capable of metabolizing unsubstituted and alkyl-substituted aromatics, including benzene, toluene, xylene, 1,3-dimethylbenzene, acenaphthene, and naphthalene, in the absence of molecular oxygen (Grbic-Galic and Vogel 1987 ; Mihelcic and Luthy 1988a , b ; Zeyer et al. 1986 ). Anaerobic degradation of aliphatic hydrocarbon has also been reported and has been linked to denitrification (Bregnard et al. 1997 ; Chayabutra and Ju 2000 ; Wilkes et al. 2002 ), sulfate reduction (Coates et al. 1997 ; Kropp et al. 2000 ; Rueter et al. 1994 ; So and Young 1999 ) and methanogenesis (Anderson and Lovley 2000 ; Zengler et al. 1999 ).
For successful biodegradation of hydrocarbon pollutants, nutrients are very important ingredients, especially nitrogen, phosphorus, and in some cases iron (Cooney 1984 ). Some of these nutrients could become limiting factors, thus affecting the biodegradation processes. Atlas ( 1985 ) reported that when a major oil spill occurred in marine and freshwater environments, the supply of carbon was significantly increased and the availability of nitrogen and phosphorus generally became the limiting factor for oil degradation. To maintain an adequate microbial population, nutrients need to be supplemented with time in sufficient quantities in the aquifer. Depending on the constituents and microbes involved in the biodegradation process, carbon:nitrogen:phosphorus ratios necessary for enhancing biodegradation fall in the range of 100:10:1 to 100:1:0.5. To avoid overapplication of nitrogen and phosphorus, it is important to understand how much carbon can be metabolized based on oxygen-limiting conditions. Nitrogen and phosphorus should be added based on the amount of carbon that can be metabolized at any given time compared to the total average concentration of carbon (i.e., petroleum contamination) in the subsurface. For example, in a full scale operation to treat the area, a net 0.6 lb per hour of pure oxygen is introduced and is assumed to be completely utilized; during aerobic microbial activity approximately 0.17 lb per hour (4 lb per day) of hydrocarbon is theoretically oxidized by microbes (using a 3.5:1oxygen:hydrocarbon stoichiometric ratio). Then, using the theoretically optimal ratio range of 100:10:1 to 100:1:0.5 C:N:P for this example, 0.4–0.04 lb per day of nitrogen and 0.04–0.02 lb per day of phosphorus may need to be added to the treatment area to keep up with the estimated carbon metabolism rate ( http://www.epa.gov/oust/pubs/tum_ch12.pdf ). In nutrient limited wastewaters, a BOD:N:P ratio of 100:5:1 is often used as a benchmark for nutrient addition. A study in a simulated aerated lagoon was performed by Slade et al. ( 2011 ) to check the effect of nitrogen at different concentration levels while keeping the phosphorus concentration constant (BOD:N of 100:0; 100:1.3; 100:1.8; 100:2.7 and 100:4.9). A synthetic wastewater was prepared in which methanol, glucose and acetate acted as the combined carbon source, ammonium chloride was used for a nitrogen source, and dipotassium hydrogen phosphate was used as a phosphorus source. When nitrogen was not supplemented, dispersed growth was observed. Increased filamentous growth was observed by increasing nitrogen levels, with a marked change in filamentous species occurring between a BOD:N ratio of 100:1.8 and 100:2.7. Nitrogen fixation occurred at a BOD:N ratio of 100:0; 100:1.3 and 100:1.8, with nitrogen loss at BOD:N ratios of 100:2.7 and 100:4.9. At a BOD:N ratio of 100:4.9, ammonium discharge was significantly greater (1.8 mg/L) than at the lower nitrogen levels (0.04–0.18 mg/L). Phosphorus behavior was more variable, however significantly more phosphorus was discharged at the lowest nitrogen level than at the highest ( p < 0.05). Based on readily available nitrogen, the BOD:N ratio at which nitrogen fixation no longer occurred was around 100:1.9. In marine environments, it was found to be more pronounced due to low levels of nitrogen and phosphorous in seawater (Floodgate 1984 ). Freshwater wetlands are typically considered to be nutrient deficient due to heavy demands of nutrients by the plants (Mitsch and Gosselink 1993 ). Therefore, additions of nutrients were necessary to enhance the biodegradation of oil pollutants (Choi et al. 2002 ; Kim et al. 2005 ). On the other hand, excessive nutrient concentrations can also inhibit the biodegradation activity (Chaillan et al. 2006 ). Several authors have reported the negative effects of high NPK levels on the biodegradation of hydrocarbons (Oudot et al. 1998 ), especially on aromatics (Carmichael and Pfaender 1997 ). An experiment was set up by Oudot et al. ( 1998 ) in the mid-tide sediments of an estuarine environment in the bay of Brest, France to study the influence of the addition of a slow-release fertilizer on the biodegradation rate of crude oil for 9 months. The weathering of total oil and fractions was monitored relative to the internal conservative biomarker 17α(H), 21β(H)-30-norhopane by computerized capillary gas chromatography. The results of biodegradation rates for total oil, aliphatics, cycloalkanes and aromatics were respectively 40 ± 7, 83 ± 6, 49 ± 10 and 55 ± 18 %. The resins and asphaltenes were not degraded. There was no significant difference in biodegradation rates observed between fertilized and non-fertilized plots, which was attributed to the high background level of N and P. It was estimated that if background level of N in the interstitial pore water of the sediment is ≥100 μmoles/liter then bioremediation through fertilization may be of limited use only. Another study on biodegradation of PAH was carried out by Carmichael and Pfaender ( 1997 ) to examine the effects of several bioremediation stimulants, including potential metabolism pathway inducers, inorganic/organic nutrients, and surfactants on the metabolism of phenanthrene and pyrene, as well as the population dynamics of PAH-degrading microorganisms in five soils with differing background PAH concentrations, exposure histories and physical properties. Increases in the populations of heterotrophic microorganisms was observed with many of the supplements, as measured by plate counts, but no increase in the populations of phenanthrene-degrading microorganisms was observed, as measured by the [ 14 C]-PAH mineralization MPN analysis or cellular incorporation of [ 14 C]-PAH. The results suggested that the PAH-degrading community at each site may be unique in their response to materials added in an attempt to stimulate PAH degradation. The characteristics of the site, including exposure history, soil type, and temporal variation may all influence their response. The effectiveness of fertilizers for the crude oil bioremediation in subarctic intertidal sediments was studied by Pelletier et al ( 2004 ). The use of poultry manure as organic fertilizer in contaminated soil was also reported (Okolo et al. 2005 ), and biodegradation was found to be enhanced in the presence of poultry manure alone. Maki et al. ( 2005 ) reported that photo-oxidation increased the biodegradability of petroleum hydrocarbons by increasing their bioavailability and thus enhancing microbial activities. Nitrogen and phosphorus may also be limiting in soils, and the acceleration of the biodegradation of crude oil or gasoline in soil and groundwater by the addition of urea-phosphate, N-P-K fertilizers, and ammonium and phosphate salts has been demonstrated in several studies (Dibble and Bartha 1979 ; Jamison et al. 1975 ; Jobson et al. 1974 ; Verstraete et al. 1976 ). Other investigators observed no increase in biodegradation rates (Lehtomaki and Niemela 1975 ) or an increase only after a delay of several months to a year (Odu 1978 ), (Raymond et al. 1976 ) when fertilizer amendments were used. These seemingly contradictory results have been attributed by Bossert and Bartha ( 1984 ) to the variable and complex composition of soils and to other factors such as nitrogen reserves and the presence of nitrogen-fixing bacteria.
Acidity or alkalinity
The acidity and alkalinity (PH) is also a very important factor in the biodegradation of petroleum hydrocarbon. In contrast to most aquatic ecosystems, soil pH can be highly variable, ranging from 2.5 in mine spoils to 11.0 in alkaline deserts (Bossert and Bartha 1984 ). Most heterotrophic bacteria and fungi prefer a pH near neutral, with fungi being more tolerant of acidic conditions (Atlas 1988 ). Extremes in pH, as can be observed in some soils, would therefore be expected to have a negative influence on the ability of microbial populations to degrade hydrocarbons. Verstraete et al. ( 1976 ) reported a near doubling of biodegradation rates of gasoline in an acidic (pH 4.5) soil by adjusting the pH to 7.4. Rates dropped significantly, however, when the pH was further raised to 8.5. Similarly, Dibble and Bartha ( 1979 ) observed an optimal pH of 7.8 in the range 5.0 to 7.8 for the mineralization of oily sludge in soil. The pH of sediments in special environments such as salt marshes may be as low as 5.0 in some cases (Patrick and DeLaune 1977 ). Hambrick et al. ( 1980 ) found the rates of microbial mineralization of octadecane and napthalene to be depressed at this pH compared with pH 6.5. Octadecane mineralization rates increased further when the pH was raised from 6.5 to 8.0, whereas naphthalene mineralization rates did not.
Soil moisture
Soil moisture is another important parameter in determining the rate of biodegradation of petroleum compounds. Microbes live in the interstitial water of soil pores and thus,a lower amount of water means a smaller number of microbes are present and the removal rate through biodegradation is slower (Dibble and Bartha 1976 ). In terrestrial ecosystems, hydrocarbon biodegradation may be limited by the availability of water for microbial growth and metabolism. Water potential (a w ) of soils can range from 0.0 to 0.99, in contrast to aquatic environments, in which water activity is stable at a value near 0.98 (Bossert and Bartha 1984 ). Atlas ( 1981 ) has suggested that tarballs deposited on beaches may represent another situation in which available water limits hydrocarbon biodegradation. Dibble and Bartha ( 1979 ) reported optimal rates of biodegradation of oil sludge in soil to be at 30–90 % water saturation. A laboratory study was conducted by Dibble and Bartha ( 1979 ) with the aim of evaluating and optimizing the environmental parameters of “landfarming,” i.e., the disposal by biodegradation in soil of oily sludges generated in the refining of crude oil and related operations. Oil sludge biodegradation was optimal at a soil water holding capacity of 30–90 %, a pH of 7.5–7.8, C:N and C:P ratios of 60:1 and 800:1, respectively, and a temperature of 20° C or above. An application rate of 5 % (wt/wt) oil sludge hydrocarbon to the soil (100,000 l/hectare) achieved a good compromise between high biodegradation rates and efficient land use, and resulted in the best overall biodegradation rate of all hydrocarbon classes. Two 100,000-liter/hectare (255 barrels per acre) or four 50,000-liter/hectare oil sludge hydrocarbon applications per growing season seem appropriate for most temperate zone disposal sites in their study. The failure to observe inhibition of degradation at the lower values was ascribed to a hydrocarbon-mediated reduction in the water holding capacity of the soil.
Bioavailability
Bioavailability is a key factor in the efficient biodegradation of pollutants. It is the amount of a substance that is physiochemically accessible to microorganisms. The application of external nonionic surfactant, e.g., the main components of oil spill dispersants, influences the alkane degradation rate (Bruheim and Eimhjelle 1998 ; Rahman et al. 2003 ). The use of surfactants in situations of oil contamination may have a stimulatory, inhibitory or neutral effect on the bacterial degradation of the oil components (Liu et al. 1995 ). Chemical dispersants caused ecological damage after application for the abatement of spilled oil in marine ecosystems (Smith 1968 ), while biosurfactants from soil or freshwater microorganisms are less toxic and partially biodegradable (Poremba et al. 1991 ). Commercially available surfactants, both ionic and nonionic in nature (Laha and Luthy 1992 ; Thai and Maier 1992 ; Pennell et al. 1993 ), as well as biosurfactants and biosurfactant-producing bacteria have been investigated for their ability to increase bioavailability (Van Dyke et al. 1991 ; Volkering et al. 1993 ; Miller 1994 ). Physical disruption of soil aggregates using sonication has been reported to increase biodegradation rates effectively in a landfarm experiment. Weissenfel et al. ( 1992 ) demonstrated that soil constituents have significant impact on the bioavailability of contaminants.
Salinity plays an important role in the degradation of hydrocarbons by microorganisms. The naphthalene mineralization rate and salinity in sediments of the Hudson river were dependent upon the ambient salinity regime, with estuarine sites exhibiting a lack of inhibition of mineralization over a wider range of salinities than was the case for the less saline upstream site (Kerr and Capone). In a study of hypersaline salt evaporation ponds, Ward and Brock ( 1978b ) showed that rates of hydrocarbon metabolism decreased with increasing salinity in the range of 3.3–28.4 % and attributed the results to a general reduction in microbial metabolic rates.
Pressure is another important factor in the biodegradation of hydrocarbons. The degradation of tetradecane, hexadecane, and a mixed hydrocarbon substrate by a mixed culture of deep-sea sediment bacteria was monitored at 1 atm (ca. 101 kPa) and 495 or 500 atm (ca. 50,140 or 50,650 kPa) by Schwarz et al. ( 1974 ; 1975 ). Colwell and Walker ( 1977 ) have suggested that oil which reaches the deep ocean environment will be degraded very slowly by microbial populations and consequently, that certain recalcitrant fractions of the oil could persist for years or decades.
Microbial degradation of petroleum hydrocarbon
In 1946, Claude E. ZoBell recognized that many microorganisms have the ability to utilize hydrocarbons as their sole source of carbon and energy. Hydrocarbons in the environment are biodegraded primarily by bacteria and fungi. Although ubiquitous in terrestrial (Jones and Edington 1968 ; Atlas et al. 1980 ) and aquatic (Mulkins-Phillips and Stewart 1974 ; Buckley et al. 1976 ; Ward and Brock 1976 ) ecosystems, the fraction of the total heterotrophic community represented by hydrocarbon-utilizing bacteria and fungi is highly variable, with reported frequencies ranging from 6 % (Jones et al. 1970 ) to 82 % (Pinholt et al. 1979 ) for soil fungi, 0.13 % (Jones et al. 1970 ) to 50 % (Pinholt et al. 1979 ) for soil bacteria, and 0.003 % (Hollaway et al. 1980 ) to 100 % (Mulkins-Phillips and Stewart 1974 ) for marine bacteria. Both bacteria and fungi are relatively plentiful in soil, and members of both groups contribute to the biodegradation of hydrocarbons (Bossert and Bartha 1984 ). Hydrocarbon-utilizing bacteria (Jensen 1975 ; Pinholt et al. 1979 ; Atlas et al. 1980 ) and fungi (Llanos and Kjoller 1976 ; Pinholt et al. 1979 ; Atlas et al. 1980 ) are readily isolated from soil and the application of oil or oily wastes to soil results in increased numbers of bacteria and fungi (Jensen 1975 ; Llanos and Kjoller 1976 ; Pinholt et al. 1979 ). In the only published comparative study of hydrocarbon degradation by bacteria and fungi in soil, Song et al. ( 1986 ) observed that 82 % of n-hexadecane mineralization in a sandy loam was attributed to bacteria and only 13 % was attributed to fungi. Individual organisms can metabolize only a limited range of hydrocarbon substrates (Britton 1984 ), so that assemblages of mixed populations with overall broad enzymatic capacities are required to degrade complex mixtures of hydrocarbons such as crude oil in soil (Bossert and Bartha 1984 ), freshwater (Cooney 1984 ), and marine (Atlas 1981 ; Floodgate 1984 ) environments. The ability to degrade and/or utilize hydrocarbon substrates is exhibited by a wide variety of bacterial and fungal genera. Floodgate ( 1984 ) lists 25 genera of hydrocarbon-degrading bacteria and 27 genera of hydrocarbon-degrading fungi which have been isolated from the marine environment; a similar compilation by Bossert and Bartha ( 1984 ) for soil isolates includes 22 genera of bacteria and 31 genera of fungi. Based on the number of published reports, the most important hydrocarbon-degrading bacteria in both marine and soil environments are Achromobacter, Acinetobacter, Alcaligenes, Arthrobacter, Bacillus , Flavobacterium, Nocardia and Pseudomonas sp. and the coryneforms; the importance of hydrocarbon-degrading strains of the salt-requiring Vibrio sp. is limited to the marine environment. Austin et al. ( 1977 ), in a numerical taxonomy study of petroleum-degrading bacteria from Chesapeake Bay water and sediment, found that Pseudomonas, Micrococcus, and Nocardia sp., members of the family Enterobacteriaceae, Actinomycetes, and coryneforms made up 95 % of the isolates. Among the fungi, Aureobasidium, Candida, Rhodotorula and Sporobolomyces sp. are the most common marine isolates and Trichoderma and Mortierella sp. are the most common soil isolates. Hydrocarbon-degrading Aspergillus and Penicillium sp. have been frequently isolated from both environments. Based on the work of Kirk and Gordon ( 1988 ), the truly marine, beach-adapted genera Corollospora, Dendryphiella, Lulworthia, and Varicosporina should be added to the list. Fungal genera, namely Amorphoteca , Neosartorya , Talaromyces, and Graphium, and yeast genera, namely Candida, Yarrowia, and Pichia , were isolated from petroleum-contaminated soil and proved to be potential organisms for hydrocarbon degradation (Chaillan et al. 2004 ). Singh ( 2006 ) also reported a group of terrestrial fungi, namely Aspergillus , Cephalosporium, and Pencillium, which were also found to be potential degraders of crude oil hydrocarbons. The yeast species, namely Candida lipolytica, Rhodotorula mucilaginosa, Geotrichum sp. and Trichosporon mucoides , when isolated from contaminated water were noted to degrade petroleum compounds (Bogusławska-Was and Dabrowski 2001 ). Walker and Colwell ( 1974 ) reported that Chesapeake Bay bacteria accounted for all of the utilization of model petroleum at 0 and 5 °C, while at 10 °C, yeasts but not filamentous fungi contributed to the utilization of the petroleum. A hydrocarbon-degrading fungus, Cladosporium resinae, was responsible for 20–40 % of the degradation of petroleum when added to the inoculum. Algae and protozoa are important members of the microbial community in both aquatic and terrestrial ecosystems, but the extent of their involvement in hydrocarbon biodegradation is largely unknown. Walker et al. ( 1975 ) isolated an alga, Prototheca zopfii , which was capable of utilizing crude oil and a mixed-hydrocarbon substrate and exhibited extensive degradation of n- and isoalkanes, as well as aromatic hydrocarbons. Cerniglia et al. ( 1980 ) observed that nine cyanobacteria, five green algae, one red alga, one brown alga, and two diatoms could oxidize naphthalene. Protozoa, by contrast, have not been shown to utilize hydrocarbons. Rogerson and Berger ( 1981 ) found no direct utilization of crude oil by protozoa cultured on hydrocarbon-utilizing yeasts and bacteria. Overall, the limited available evidence does not appear to suggest an ecologically significant role for algae and protozoa in the degradation of hydrocarbons in the environment (O’Brien and Dixon 1976 ; Bossert and Bartha 1984 ). Some hydrocarbon-degrading microorganisms are listed in Table 3 .
Mechanism of petroleum hydrocarbon degradation
The most rapid and complete degradation of the majority of organic pollutants is brought about under aerobic conditions. The initial intracellular attack of organic pollutants is an oxidative process and the activation as well as incorporation of oxygen is the enzymatic key reaction catalyzed by oxygenases and peroxidases. Peripheral degradation pathways convert organic pollutants step by step into intermediates of the central intermediary metabolism; for example, the tricarboxylic acid cycle. Biosynthesis of cell biomass occurs from the central precursor metabolites, such as acetyl-CoA, succinate, and pyruvate. Sugars required for various biosyntheses and growth are synthesized by gluconeogenesis. The degradation of petroleum hydrocarbons can be mediated by specific enzyme systems. Other mechanisms involved are (1) attachment of microbial cells to the substrates and (2) production of biosurfactants (Hommel 1990 ). The uptake mechanism linked to the attachment of cells to oil droplets is still unknown but production of biosurfactants has been well studied. Biosurfactants can act as emulsifying agents by decreasing the surface tension and forming micelles. The microdroplets encapsulated in the hydrophobic microbial cell surface are taken inside and degraded. Biosurfactants produced by some microorganisms are listed in Table 5 , below. Cameotra and Singh ( 2008 ) reported bioremediation of oil sludge using biosurfactants; the microbial consortium used in their study consisted of two isolates of Pseudomonas aeruginosa and one isolate of Rhodococcus erythropolis from oily sludge contaminated soil. Ninety percent of the hydrocarbons were degraded by the consortium after 6 weeks in liquid culture. The objective of their study was to test the ability of the consortium to degrade sludge hydrocarbons in two separate field trials and to assess the effect of two additives—a nutrient mixture and a crude biosurfactant preparation—on the efficiency of the process. The biosurfactant used was identified as a mixture of 11 rhamnolipid congeners and was produced by a consortium member. The observation of their study showed that 91 % of the hydrocarbon content from the soil contaminated with 1 % (v/v) crude oil sludge was degraded by the consortium after 5 weeks. About 91–95 % depletion of the hydrocarbon content was reported with the separate use of any one additive along with the consortium after 4 weeks. The crude biosurfactant preparation proved to be a more effective enhancer of degradation. When both the additives were combined with the consortium, more than 98 % hydrocarbon depletion was achieved. This observation suggested that hydrocarbon remediation could be successfully achieved through the use of a crude biosurfactant (Das and Chandran 2011 ). Biosurfactants produced by some microorganisms are listed in Table 5 .
Role of enzymes in degradation of hydrocarbons
Cytochrome P450 alkane hydroxylases constitute a super family of ubiquitous heme-thiolate monooxygenases, which play an important role in the microbial degradation of oil, chlorinated hydrocarbons, fuel additives, and many other compounds (Van Beilen and Funhoff 2007 ). Depending on the chain length, enzyme systems are required to introduce oxygen in the substrate to initiate biodegradation. Higher eukaryotes generally contain several different P450 families that consist of large number of individual P450 forms that may contribute as an ensemble of isoforms to the metabolic conversion of a given substrate. In microorganisms such as P450, multiplicity can only be found in a few species (Zimmer et al. 1996 ). The cytochrome P450 enzyme system was found to be involved in the biodegradation of petroleum hydrocarbons (Das and Chandran 2011 ). The capability of several yeast species to use n-alkanes and other aliphatic hydrocarbons as a sole source of carbon and energy is mediated by the existence of multiple microsomal Cytochrome P450 forms. These cytochrome P450 enzymes had been isolated from yeast species such as Candida maltosa , Candida tropicalis , and Candida apicola (Scheuer et al. 1998 ). The diversity of alkaneoxygenase systems in prokaryotes and eukaryotes that are actively participating in the degradation of alkanes under aerobic conditions, such as Cytochrome P450 enzymes, integral membrane di-iron alkane hydroxylases (e.g., alkB ), soluble di-iron methane monooxygenases, and membrane-bound copper containing methane monooxygenases, have been discussed by Van Beilen and Funhoff ( 2005 ). Enzymes and Biosurfactants involved in the biodegradation of petroleum hydrocarbon are listed in Tables 4 and 5 .
Genetic aspects of petroleum hydrocarbon degradation
Genetic factors play an important role in conferring biodegradation potentials on microorganisms. Plasmid DNA plays an important role in genetic adaptation, as it represents a highly mobile form of DNA which can be transferred via conjugation or transformation and can impart novel phenotypes, including the ability to transmit hydrocarbon-oxidation to a recipient organism. The pathways for the metabolism of naphthalene, salicylat, camphor, octane, xylene and toluene have been encoded on plasmids in Pseudomonas sp. (Chakrabarty 1976 ). The ability to degrade more recalcitrant components of petroleum, such as the aromatic fractions, is generally plasmid mediated (Cerniglia 1984b ). The observed increase in the study of genetics paralleled the development of advances in molecular biology, particularly the application of recombinant DNA technology (Singer and Finnerty 1984b ), gene probes (Barriault and Sylvestre 1993 ), and polymerase chain reaction (PCR) technology. Many bacterial catabolic pathways are specified by conjugative plasmids (Frantz and Chakrabarty 1986 ). These plasmids are readily transferred laterally into new host bacteria, thereby enhancing the metabolic potential of other members of an ecosystem (Okoh 2006 ).
Phytoremediation
Phytoremediation is an emerging technology that uses plants in the cleanup of environmental pollution, with hydrocarbons and other hazardous substances. Advantages of using phytoremediation include cost-effectiveness, aesthetic advantages, and long-term applicability. The Alabama Department of Environmental Management studied a 1,500-cubic-yard soil site, of which 70 % of the baseline samples contained over 100 ppm of total petroleum hydrocarbon (TPH). After 1 year of vegetative cover, approximately 83 % of the samples were found to contain less than 10-ppm TPH. Removal of total petroleum hydrocarbon (TPH) at several field sites contaminated with crude oil, diesel fuel, or petroleum refinery wastes at initial TPH concentrations of 1,700–16,000 mg/kg were also investigated (Hecht and Badiane 1998 ; Nedunuri et al. 2000 ). Plant growth was found to vary depending upon the species. The presence of some species led to greater TPH disappearance than with other species or compared to unvegetated soil. Among tropical plants tested for use in the Pacific Islands, three coastal trees, kou ( Cordia subcordata ), milo ( Thespesia populnea ), and kiawe ( Prosopis pallida ), and the native shrub beach naupaka ( Scaevola serica ) were found to tolerate field conditions and facilitated the cleanup of soils contaminated with diesel fuel (U.S. Army Corps of Engineers 2003 ). Microbial degradation in the rhizosphere might be the most significant mechanism for removal of diesel range organics in vegetated contaminated soils (Miya and Firestone 2001 ). This occurs because contaminants such as PAHs are highly hydrophobic, and their sorption to soil decreases their bioavailability for plant uptake and phytotransformation (Das and Chandran 2011 ).
Future prospects
The bioremediation studies after the Exxon Valdez oil spill demonstrated the promise of this technology for cleaning up a petroleum-contaminated site (NOAA Technical Memorandum 1997 ). Bioremediation was more extensive in 1990, with 378 of the 587 shoreline segments treated that year receiving bioremediation application (Chianelli et al. 1991 ). To determine the extent of the natural removal of oil, the National Oceanic and Atmospheric Administration (NOAA) conducted monitoring programs which determined that removal of surface oil from exposed shorelines was about 90%; for sheltered shorelines and those with intermittent energy, the removal rates were 70%. For subsurface oil, the removal rate was approximately 55%, though this varied with the depth of the subsurface oil impacts (Michel et al. 1991 ). It has been estimated that a beach damaged by an oil spill could be returned to its natural condition in as little as 2–5 years with bioremediation treatment; otherwise it would take 10 years or more to reach this condition. Ten years after the Exxon Valdez accident, oil residues still remain below the surface in the spill-affected bays where contamination problems persist and negatively impact fishery activities. This fact clearly indicates the necessity for further development of bioremediation technologies to manage marine and terrestrial oil pollution. The microorganisms which play major roles in bioremediation, and their nutrient requirements (carbon, nitrogen, phosphorous, etc.) and environmental requirements (oxygen or an alternative electron acceptor, temperature, redox potential, salinity, pH, etc.) should be further researched and confirmed. Such knowledge would allow the manipulation of environmental factors that may limit or prevent the biodegradation of petroleum in environment. Undertaking bioremediation cannot be done without the agreement of local communities. Their concerns about bioremediation, a relatively new technology, should be mitigated by scientists who can explain to the local people the results of hydrocarbon contamination tests and microorganism tests, especially in regard to risk assessment. The identification of the Alcanivorax group as a major population that arose during a bioremediation treatment would be useful information to help convince local residents that the propagation of harmful microorganisms would not occur during bioremediation (Harayama et al. 1999 ).
Bioremediation is a very popular and promising technology for the remediation of environments contaminated with petroleum hydrocarbon. Petroleum hydrocarbon contamination causes various hazardous effects on ecosystems and humans. Physical and chemical methods used for the treatment are not that much more efficient and are very costly compared to bioremediation, which is less expensive and more efficient. Various physical and chemical factors, such as temperature, oxygen, pH, nutrients etc. and the type of contaminants, metabolic capabilities of the indigenous microbial population, and type of plant species used in phytoremediation affect the degradation rate of petroleum hydrocarbon. Microbial degradation of petroleum hydrocarbon is accomplished by a diversity of microorganisms because they have enzyme systems to degrade and utilize different hydrocarbons as a source of carbon and energy. Bacteria are the most powerful degrading agent compared to fungi, algae and protozoa. Genetically engineered microorganisms (GEMs) and phytoremediation also play an important role in the remediation of petroleum hydrocarbon from environment. Several GEMs have been successfully constructed and experimentally proven to have a higher degradative capability and utility for bioremediation under laboratory conditions (Furukawa 2003 ; Pieper and Reineke 2000 ; Lovley 2003 ). In spite of such studies, applying GEMs in situ has been limited by the risks associated with horizontal gene transfer and uncontrolled proliferation of the introduced GEMs (Velkov 2001 ; Boopathy 2000 ). Austria, France, Greece, Hungary, Germany and Luxembourg have all banned the cultivation of genetically modified organisms (GMOs). Angola has banned imports of all GMO produce, except for food aid provided it was milled; Zambia banned import of all GMOs as well. Therefore, from this present review we conclude that the microbial degradation of petroleum hydrocarbons can be considered as a key component in the cleanup strategy for petroleum hydrocarbon remediation.
Alexander M (1999) Biodegradation and bioremediation. Academic, San Diego
Google Scholar
Alloway BJ and Ayres DC (1993) Organic pollutants. In: Chemical principles of environmental pollution. 1st edition. Chapman and Hall Publishers, India, p 201
Alvarez PJJ, Vogel TM (1991) Substrate interactions of benzene, toluene, and para-xylene during microbial degradation by pure cultures and mixed culture aquifer slurries. Appl Environ Microbiol 57:2981–2985
PubMed CAS Google Scholar
Anderson RT, Lovley DR (2000) Hexadecane decay by methanogenesis. Nature 404:722–723
Article PubMed CAS Google Scholar
Anonymous (1989) Mishaps cause three oil spills off U.S. Oil Gas J 87:22
April TM, Foght JM, Currah RS (2000) Hydrocarbon-degrading filamentous fungi isolated from flare pit soils in northern and western Canada. Can J Microbiol 46:38–49
Atlas RM (1975) Effects of temperature and crude oil composition on petroleum biodegradation. J Appl Microbiol 30:396–403
CAS Google Scholar
Atlas RM (1981) Microbial degradation of petroleum hydrocarbons: an environmental perspective. Microbiol Rev 45:180–209
Atlas RM (1985) Effects of hydrocarbons on micro-organisms and biodegradation. In: Engelhardt FR (ed) Arctic ecosystems in petroleum effects in the arctic environment. Elsevier, London, pp 63–99
Atlas RM (1988) Microbiology-fundamentals and applications, 2nd edn. Macmillan Publishing Co, New York, pp 352–354
Atlas RM, Bartha R (1972) Biodegradation of petroleum in seawater at low temperatures. Can J Microbial 18:1851–1855
Article CAS Google Scholar
Atlas RM, Schofield EA, Morelli FA, Cameron RE (1976) Interactions of microorganisms and petroleum in the Arctic. Environ Pollut 10:35–44
Article Google Scholar
Atlas RM, Horowitz A, Busdosh M (1978) Prudhoe crude oil in Arctic marine ice, water, and sediment ecosystems: degradation and interactions with microbial and benthic communities. J Fish Res Board Can 35:585–590
Atlas RM, Sexstone A, Gustin P, Miller O, Linkins P, Everett K (1980) Biodegradation of crude oil by tundra soil microorganisms. In: Oxley TA, Becker G and Allsop D (eds) Proceedings of the 4th International Biodeterioration. Symposium. Pitman Publishing Ltd., London, p 21–28
Austin B, Calomiris JJ, Walker JD, Colwell RR (1977) Numerical taxonomy and ecology of petroleum-degrading bacteria. Appl Environ Microbiol 34:60–68
Bahram A (1969) Solubility of light hydrocarbons in water under high pressures. Ph.D thesis. University of Oklahoma
Bailey NJL, Jobson AM, Rogers MA (1973) Bacterial degradation of crude oil: comparison of field and experimental data. Chem Geol 11:203–221
Barriault D, Sylvestre M (1993) Factors affecting PCB degradation by an implanted bacterial strain in soil microcosms. Can J Microb 39:594–602
Bartha R, Bossert I (1984) The treatment and disposal of petroleum wastes. In: Atlas RM (ed) Petroleum microbiology. Macmillan, New York, pp 553–578
Bhupathiraju VK, Krauter P, Holman HYN, Conrad ME, Daley PF, TempletonAS HJR, Hernandez M, Alvarez-Cohen L (2002) Assessment of in-situ bioremediation at a refinery waste-contaminated site and an aviation gasoline contaminated site. Biodegradation 13:79–90
Bidoia ED, Montagnolli RN, and Lopes PRM (2010) Microbial biodegradation potential of hydrocarbons evaluated by colorimetric technique: a case study. Appl Microbiol Microb Biotechnol A. Mendas-Vilas (ed)
Bogusławska-Was E, Dabrowski W (2001) The seasonal variability of yeasts and yeast-like organisms in water and bottom sediment of the Szczecin Lagoon. Int J Hyg Environ Health 203:451–458
Article PubMed Google Scholar
Boopathy R (2000) Factors limiting bioremediation technologies. Bioresour Technol 74:63–67
Bossert I, Bartha R (1984) The fate of petroleum in soil ecosystems. In: Atlas RM (ed) Petroleum microbiology. Macmillan Publishing Co, New York, pp 434–476
Boyd SA, Shelton DR (1984) Anaerobic biodegradation of chlorophenols in fresh and acclimated sludge. Appl Environ 47:272–277
Bregnard TP-A, Häner A, Höhener P, Zeyer J (1997) Anaerobic degradation of pristane in nitrate-reducing microcosms and enrichment cultures. Appl Environ Microbiol 63:2077–2081
Britton LN (1984) Microbial degradation of aliphatic hydrocarbons. In: Gibson DT (ed) Microbial degradation of organic compounds. Marcel Dekker, Inc, New York, pp 89–129
Bruheim P, Eimhjelle K (1998) Chemically emulsified crude oil as substrate for bacterial oxidation; differences in species response. Can J Micro 442:195199
Brusseau ML (1998) The impact of physical, chemical and biological factors on biodegradation. In Serra R (ed) Proceedings of the international conference on biotechnology for soil remediation: scientific bases and practical applications. C.I.P.A. S.R.L., Milan, Italy, pp 81–98
Buckley EN, Jones RB, Pfaender FK (1976) Characterization of microbial isolates from an estuarine ecosystem: relationship of hydrocarbon utilization to ambient hydrocarbon concentrations. Appl Environ Microbiol 32:232–237
Cameotra SS, Singh P (2008) Bioremediation of oil sludge using crude biosurfactants. Int Biodeterior Biodegrad 62:274–280
Caplan JA (1993) The world-wide bioremediation industry: prospects for profit. Trends Biotech 11:320–323
Carmichael LM, Pfaender FK (1997) The effect of inorganic and organic supplements on the microbial degradation of phenanthrene and pyrene in soils. Biodegradation 8(1):1–13
Cerniglia CE (1984a) Microbial degradation of polycyclic aromatic hydrocarbons. Adv Appl Microb 30:31–71
Cerniglia CE (1984b) Microbial transformation of aromatic hydrocarbons. In: Atlas RM (ed) Petroleum microbiology. Macmillan Publishing Co, New York, pp 99–128
Cerniglia CE, Gibson DT, van Baalen C (1980) Oxidation of naphthalene by cyanobacteria and microalgae. J Gen Microbiol 116:495–500
Chaillan F, Le Flèche A, Bury E, Phantavong YH, Grimont P, Saliot A, Oudot J (2004) Identification an biodegradation potential of tropical aerobic hydrocarbon degrading microorganisms. Res Microbiol 155:587–595
Chaillan F, Chaineau CH, Point V, Saliot A, Oudot J (2006) Factors inhibiting bioremediation of soil contaminated with weathered oils and drill cuttings. Environ Pollut 144:255–265
Chakrabarty AM (1976) Plasmids in pseudomonas . Ann Rev Genet 10:7–30
Chapelle FH (1999) Bioremediation of petroleum hydrocrbon–contaminated ground water: the perspectives of history and hydrology. Ground Water 37 No. 1
Chayabutra C, Ju L (2000) Degradation of n -hexadecane and its metabolites by Pseudomonas aeruginosa under micro aerobic and anaerobic denitrifying conditions. Appl Environ Microbiol 66:493–498
Chen M, Hong CS, Bush B, Rhee G-Y (1988) Anaerobic biodegradation of polychlorinated biphenyls by bacteria from Hudson River sediments. Ecotoxicol Environ Saf 16:95–105
Chianelli RR, Aczel T, Bare RE, George GN, Genowitz MW, Grossman MJ, Haith CE, Kaiser FJ, Lessard RR, Liotta R, Mastracchio RL, Minak-Bernero V, Prince RC, Robbins WK, Stiefel EI, Wilkinson JB, Hinton SM, Bragg JR, McMillen SJ, Atlas RM (1991) Bioremediation technology development and application to the alaskan spill. Oil Spill Conference Proceedings 1991, pp 549–558
Choi S-C, Kwon KK, Sohn JH, Kim S-J (2002) Evaluation of fertilizer additions to stimulate oil biodegradation in sand seashore mesocosms. J Microbiol Biotechnol 12:431–436
Coates JD, Woodward J, Allen J, Philp P, Lovley DR (1997) Anaerobic degradation of polycyclic aromatic hydrocarbons and alkanes in petroleumcontaminated marine harbour sediments. Appl Environ Microbiol 63:3589–3593
Colwell RR, Walker JD (1977) Ecological aspects of microbial degradation of petroleum in the marine environment. Crit Rev Microbiol 5:423–445
Colwell RR, Mills AL, Walker JD, Garcia-Tello P, Campos PV (1978) Microbial ecology of the Metula spill in the Straits of Magellan. J Fish Res Board Can 35:573–580
Cooney JJ (1984) The fate of petroleum pollutants in fresh water ecosystems. In: Atlas RM (ed) Petroleum microbiology. Macmillan, New York, pp 399–434
Cooney JJ, Silver SA, Beck EA (1985) Factors influencing hydrocarbon degradation in three freshwater lakes. Microb Ecol 11:127–137
Culberson OL (1951) Solubility of methane and ethane in water at temperatures of 340 °F and pressure to 10,000 psia. Ph.D. thesis, University of Texas
Das N, Chandran P (2011) Microbial degradation of petroleum hydrocarbon contaminants: an overview. SAGE-Hindawi Access to Research; Biotechnology Research International Volume 2011, 13 pages
Daverey A, Pakshirajan K (2009) Production of sophorolipids by the yeast Candida bombicola using simple and low cost fermentative media. Food Res Int 42:499–504
Delille D, Coulon F, Pelletier E (2004) Effects of temperature warming during a bioremediation study of natural and nutrient-amended hydrocarbon-contaminated sub-Antarctic soils. Cold Regions Sci Tech 40:61–70
Desai A, Pranav V (2006) Petroleum and hydrocarbon microbiology. Applied microbiology. Report of Department of Microbiology. M.S. University of baroda, Vadodara, India, p 22
Dibble JT, Bartha R (1976) The effect of iron on the biodegradation of petroleum in seawater. Appl Environ Microbiol 31:544–550
Dibble JT, Bartha R (1979) Effect of environmental parameters on the biodegradation of oil sludge. Appl Environ Microbiol 37:729–739
Fedorak PM, Westlake DWS (1981) Microbial degradation of aromatics and saturates in Prudhoe Bay crude oil as determined by glass capillary gas chromatography. Can J Microbiol 27:432–443
Floodgate G (1984) The fate of petroleum in marine ecosystems, In: Atlas RM (ed) Petroleum microbiology. Macmillion, New York, pp 355–398
Foght JM, Westlake DWS, Johnson WM, Ridgway HF (1996) Environmental gasoline-utilizing isolates and clinical isolates of Pseudomonas aeruginosa are taxonomically indistinguishable by chemotaxonomic and molecular techniques. Microbiology 142:2333–2340
Frantz B, Chakrabarty AM (1986) Degradative plasmids in pseudomonas aeruginosa. In: Sokatch JR, Ornston LN (eds) The bacteria, vol 10. Academic, New York, pp 295–32
Furukawa K (2003) ‘Super bugs’ for bioremediation. Trends Biotechnol 21:187–190
Fusey P, Oudot J (1984) Relative influence of physical removal and biodegradation in the depuration of petroleum contaminated seashore sediments. Mar Pollut Bull 15:136–141
Gibbs CF, Pugh KB, Andrews AR (1975) Quantitative studies on marine biodegradation of oil. II. Effect of temperature. Proc R Soc London, Ser B 188:83–94
Grbic-Galic D, Vogel TM (1987) Transformation of toluene and benzene by mixed methanogenic cultures. Appl Environ Microbiol 53:254–260
Gupta R, Mahapatra H (2003) Microbial biomass: an economical alternative for removal of heavy metals from waste water. Indian J Experiment Biol 41:945–966
Hagar R (1989) Huge cargo of North Slope oil spilled. Oil Gas J 87:26–27
Hambrick GA, DeLaune RD, Patrick WH (1980) Effect of estuarine sediment pH and oxidation-reduction potential on microbial hydrocarbon degradation. Appl Environ Microbiol 40:365–369
Harayama S, Kishira H, Kasai Y, Shutsubo K (1999) Petroleum biodegradation in marine environments. J Molec Microbiol Biotechnol 1:63–70
Hecht D, Badiane G (1998) Phytoremediation. New Internationalist
Hernandez A, Mellado RP, Martinez JL (1998) Metal accumulation and vanadiuminduced multidrug resistance by environmental isolates of Escherichia herdmann and Enterobacter cloacae. Appl Environ Microbiol 64:4317–4320
Hollaway SL, Faw GM, Sizemore RK (1980) The bacterial community composition of an active oil field in the Northwestern Gulf of Mexico. Mar Pollut Bull 11:153–156
Hommel RK (1990) Formation and phylogenetic role of biosurfactants. J Appl Microbiol 89:158–119
http://esc.syrres.com/efdb/chemfate.htm
http://www.epa.gov/oust/pubs/tum_ch12.pdf
Huddleston RL and Cresswell LW (1976) Environmental and nutritional constraints of microbial hydrocarbon utilization in the soil. In: Proceedings of the 1975 Engineering Foundation Conference: the role of microorganisms in the recovery of oil. National Science Foundation, Washington, D.C., p 71–72
Iida T, Sumita T, Ohta A, Takagi M (2000) The cytochrome P450ALKmultigene family of an n-alkane-assimilating yeast, Yarrowia lipolytica : cloning and characterization of genes coding for new CYP52 family members. Yeast 16:1077–1108
Ilori MO, Amobi CJ, Odocha AC (2005) Factors affecting biosurfactant production by oil degrading Aeromonas spp. isolated from a tropical environment. Chemosphere 61:985–992
International Agency for Research on Cancer (IARC) (2000) Monographs on the valuation of the carcinogenic risk of some industrial chemicals to humans. Lyon, France
Jamison VM, Raymond RL, Hudson JO Jr (1975) Biodegradation of high-octane gasoline in groundwater. Dev Ind Microbiol 16:305–312
Jan B, Beilen V, Neuenschwunder M, Suits THM, Roth C, Balada SB, Witholt B (2003) Rubredoxins involved in alkane degradation. J Bacteriol 184:1722–1732
Jensen V (1975) Bacterial flora of soil after application of oily waste. Oikos 26:152–158
Jobson A, Cook FD, Westlake DWS (1972) Microbial utilization of crude oil. Appl Microbiol 23:1082–1089
Jobson A, McLaughlin M, Cook FD, Westlake DWS (1974) Effect of amendments on the microbial utilization of oil applied to soil. Appl Microbiol 27:166–171
Jones JG, Edington MA (1968) An ecological survey of hydrocarbon-oxidizing micro-organisms. J Gen Microbiol 52:381–390
Jones JG, Knight M, Byron JA (1970) Effect of gross pollution by kerosine hydrocarbons on the microflora of a moorland soil. Nature 227:1166
Jones DM, Douglas AG, Parkes RJ, Taylor J, Giger W, Schaffner C (1983) The recognition of biodegraded petroleum- derived aromatic hydrocarbons in recent marine sediments. Mar Pollut Bull 14:103–108
Kim S-J, Choi DH, Sim DS, Oh Y-S (2005) Evaluation of bioremediation effectiveness on crude oil-contaminated sand. Chemosphere 59:845–852
Kirk PW, Gordon AS (1988) Hydrocarbon degradation by filamentous marine higher fungi. Mycologia 80:776–782
Kobayashi R (1951) Vapor-liquid equilibria in binary hydrocarbon-water system. Ph.D. Thesis, University of Michigan
Kropp KG, Davidova IA, Suflita JM (2000) Anaerobic oxidation of n-dodecane by an addition reaction in a sulfatereducing bacterial enrichment culture. Appl Environ Microbiol 66:5393–5398
Kumar M, Leon V, De Sisto MA, Ilzins OA, Luis L (2008) Biosurfactant production and hydrocarbondegradation by halotolerant and thermotolerant Pseudomonas sp. World J Microbiol Biotechnol 24:1047–1057
Kvenvolden KA, Cooper CK (2003) Natural seepage of crude oil into the marine environment. Geo-Mar Lett 23:140–146
Laha S, Luthy RG (1992) Effects of nonionic surfactants on the solubilization and mineralization of phenanthrene in soil-water system. Biotechnol Bioeng 14:1367–1380
Leeson A, Hinchee RE (1997) Soil bioventing, principles and practice. CRC, Lewis Publishers, Boca Raton
Lehtomaki M, Niemela S (1975) Improving microbial degradation of oil in soil. Ambio 4:126–129
Litchfield JH, Clark LC (1973) Bacterial activity in ground waters containing petroleum products. American Petroleum Institute Publication No.4211. Washington, D.C., API
Liu Z, Jacobson AM, Luthy RG (1995) Biodegradation of naphthalene in aqueous nonionic surfactant system. Appl Environ Microb 61:145–151
Llanos C, Kjoller A (1976) Changes in the flora of soil fungi following oil waste application. Oikos 27:377–382
Lovley DR (2003) cleaning up with genomics: applying molecular biology to bioremediation. Nat Rev Microbiol 1:35–44
Maeng JHO, Sakai Y, Tani Y, Kato N (1996) Isolation and characterization of a novel oxygenase that catalyzes the first step of n-alkane oxidation in Acinetobacter sp. strain M-1. J Bacteriol 178:3695–3700
Mahmound A, Aziza Y, Abdeltif A, Rachida M (2008) Biosurfactant production by Bacillus strain injected in the petroleum reservoirs. J Ind Microbiol Biotechnol 35:1303–1306
Maki H, Sasaki T, Haramaya S (2005) Photooxidation of biodegradable crude oil and toxicity of the photooxidized products. Chemosphere 44:1145–1151
Manning FS, Thompson RE (1991) Oilfield processing.Volume 2: crude oil. Pennwell Publishing Company, Tulsa, Oklahoma
Margesin R, Schinner F (2001) Bioremediation (natural attenuation and biostimulation) of diesel-oil-contaminated soil in an alpine glacier skiing area. Appl Environ Microbiol 67:3127–3133
Matsumiya Y, Kubo M (2007) Bioprocess handbook, 1st edn. NTS Publishing, Tokyo
McDonald IR, Miguez CB, Rogge G, Bourque D, Wendlandt KD, Groleau D, Murrell JC (2006) Diversity of solublemethane monooxygenase-containing methanotrophs isolated from polluted environments. FEMS Microbiol Lett 255:225–232
Mesarch MB, Nakatsu CH, Nies L (2000) Development of catechol 2,3 –Deoxygenase-specific primers for monitoring bioremediation by competitive quantitative PCR. Appl Environ Microb 66:678–690
Michel J, Hayes MO, Sexton WJ, Gibeaut JC, Henry C (1991) Trends in natural removal of the Exxon Valdez oil spill in Prince William Sound from September 1989 to May 1990. Oil Spill Conference Proceedings 1991, pp 181–187
Mihelcic JR, Luthy RG (1988a) Microbial degradation of acenaphthene and naphthalene under denitrification conditions in soil-water systems. Appl Environ Microbiol 54:1188–1198
Mihelcic JR, Luthy RG (1988b) Degradation of polycyclic aromatic hydrocarbons under various redox conditions in soil-water systems. Appl Environ Microbiol 54:1182–1187
Milic JS, Vladimir PB, Mila VI, Samira AMA, Gordana DG-C, Miroslav MV (2009) Bioremediation of soil heavily contaminated with crude oil and its products: composition of the microbial consortium. J Serb Chem Soc 74:455–460
Miller R (1994) Surfactant enhanced bioavailability of slightly soluble organic compounds. In: Skipper H (ed) Bioremediation-science and application. Soil Science Society of American Publications, Madison
Mitsch WJ, Gosselink JG (1993) Wetlands, 2nd edn. John Wiley publ, New York
Miya RK, Firestone MK (2001) Enhanced phenanthrene biodegradation in soil by slender oat root exudates and root debris. J Environ Qual 30:1911–1918
Mortelmans K, Harworth S, Lawlor T (1986) Sp Environ Mutagen 8:1
Mulkins-Phillips GJ, Stewart JE (1974) Distribution of hydrocarbon-utilizing bacteria in northwestern Atlantic waters and coastal sediments. Can J Microbiol 20:955–962
Muthuswamy K, Gopalakrishnan S, Ravi TK, Sivachidambaram P (2008) Biosurfactants: properties, commercial production and application. Curr Sci 94:736–747
Nedunuri KV, Govindaraju RS, Banks MK, Schwab AP, Chen Z (2000) Evaluation of phytoremediation for field-scale degradation of total petroleum hydrocarbons. J Environ Eng 126:483–490
O’Brien PY, Dixon PS (1976) The effects of oil and oil components on algae; a review. Br Phycol J 11:115–142
Odu CTI (1978) The effect of nutrient application and aeration on oil degradation in soil. Environ Pollut 15:235–240
Okoh AI (2006) Biodegradation alternative in the cleanup of petroleum hydrocarbon pollutants. Biotechnol Mol Biol Rev 1:38–50
Okolo JC, Amadi EN, Odu CTI (2005) Effects of soil treatments containing poultry manure on crude oil degradation in a sandy loam soil. Appl Ecol Environ Res 3:47–53
Oudot J, Merlin FX, Pinvidic P (1998) Weathering rates of oil components in a bioremediation experiment in estuarine sediments. Mar Environ Res 45:113–125
Patrick WH Jr, DeLaune RD (1977) Chemical and biological redox systems affecting nutrient availability in coastal wetlands. Geosci Man 18:131–137
Pelletier E, Delille D, Delille B (2004) Crude oil bioremediation in sub- Antarctic intertidal sediments: chemistry and toxicity of oiled residues. Mar Environ Res 57:311–327
Pennell KD, Abriola LM Jr, Weber WJ (1993) Surfactants enhanced solubilization of residual dodecane in soil columns experimental investigation. Environ Sci Technol 27:2332–2340
Perry JJ (1984) Microbial metabolism of cyclic alkanes. In: Atlas RM (ed) Petroleum microbiology. Macmillan Publishing Co, New York, pp 61–98
Pieper DH, Reineke W (2000) Engineering bacteria for bioremediation. Curr Opin Biotechnol 11:262–270
Pinholt Y, Struwe S, Kjoller A (1979) Microbial changes during oil decomposition in soil. Holarct Ecol 2:195–200
Poremba K, Gunkel W, Lang S, Wagner F (1991) Marine biosurfactants III. Toxicity testing with marine microorganisms and comparison with synthetic detergents. Z Natreforsch 46:197–203
Potter TL, Simmons KE (1998) Composition of petroleum mixtures. Volume 2 of the total petroleum hydrocarbon criteria working group series. Amherst Scientific Publishers, Anherst, Massachusetts
Rahman KSM, Rahman TJ, Kourkoutas Y, Petsas I, Marchant R, Banat IM (2003) Enhanced bioremediation of n-alkane in petroleum sludge using bacterial consortium amended with rhamnolipid and micronutrients. Bioresour Technol 90:150–168
Raymond RL, Hudson JO, Jamison VW (1976) Oil degradation in soil. Appl Environ Microbiol 31:522–535
Rogerson A, Berger J (1981) Effect of crude oil and petroleum-degrading microorganisms on the growth of freshwater and soil protozoa. J Gen Microbiol 124:53–59
Rosenberg E, Legmann R, Kushmaro A, Taube R, Adler E, Ron EZ (1992) Petroleum bioremediation –a multiphase problem. Biodegradation 3:337–350
Rueter P, Rabus R, Wilkes H, Aeckersberg F, Rainey FA, Jannasch HW, Widdel F (1994) Anaerobic oxidation of hydrocarbons in crude oil by new types of sulphate reducing bacteria. Nature 372:455–4458
Scheuer U, Zimmer T, Becher D, Schauer F, Schunck W-H (1998) Oxygenation cascade in conversion of n-alkanes to α, ω-dioic acids catalyzed by cytochrome P450 52A3. J Biol Chem 273:32528–32534
Schwarz JR, Walker JD, Colwell RR (1974) Growth of deep-sea bacteria on hydrocarbons at ambient and in situ pressure. Dev Ind Microbiol 15:239–249
Schwarz JR, Walker JD, Colwell RR (1975) Deep-sea bacteria: growth and utilization of n-hexadecane at in situ temperature and pressure. Can J Microbiol 21:682–687
Shiaris MP (1989) Seasonal biotransformation of naphthalene, phenanthrene, and benzo[a]pyrene in surficial estuarine sediments. Appl Environ Microbiol 55:1391–1399
Singer ME, Finnerty WR (1984a) Microbial metabolism of straight-chain and branched alkanes. In: Atlas RM (ed) Petroleum microbiology. Macmillan Publishing Co, New York, pp 1–60
Singer JT, Finnerty WR (1984b) Genetics of hydrocarbon-utilising microorganisms. In: Atlas RM (ed) Petroluem microbiology. Macmillan Publishing Company Inc, New York, pp 299–354
Singh H (2006) Mycoremediation: fungal bioremediation. Wiley- Interscience, NewYork
Book Google Scholar
Slade AH, Thorn GJ, Dennis MA (2011) The relationship between BOD:N ratio and wastewater treatability in a nitrogen-fixing wastewater treatment system. Water Sci Technol 63:627–632
Smith J (1968) Torrey canyon pollution and marine life. Report by the Plymouth Laboratory of the Marine Biological Association of the United Kingdom, London, Cambridge University Press, 1906
So CM, Young LY (1999) Isolation and characterization of a sulfate-reducing bacterium that anaerobically degrades alkanes. Appl Environ Microbiol 65:2969–2976
Song HG, Pedersen TA, Bartha R (1986) Hydrocarbon mineralization in soil: relative bacterial and fungal contribution. Soil Biol Biochem 18:109–111
Strausz OP, Lown EM (2003) The chemistry of Alberta oil sands, bitumens, and heavy oils. Alberta Energy Research Institute, Calgary
Strong PJ, Burgess JE (2008) Treatment methods for wine-related ad distillery wastewaters: a review. Biorem J 12:70–87
Suflita JM, Horowitz A, Shelton DR, Tiedje JM (1982) Dehalogenation: a novel pathway for the anaerobic biodegradation of haloaromatic compounds. Science 218:1115–1117
Tabatabaee A, Assadi MM, Noohi AA, Sajadian VA (2005) Isolation of biosurfactant producing bacteria from oil reservoirs. Ira J Environ Health Sci Eng 2:6–12
Tarvin D, Buswell AM (1934) The methane fermentation of organic acids and carbohydrates. J Am Chem Soc 56:1751–1755
NOAA Technical Memorandum (1997) Integrating physical and biological studies of recovery from the Exxon Valdez Oil Spill. Office of Ocean Resources Conservation and Assessment National Ocean Service National Oceanic and Atmospheric Administration U.S. Department of Commerce. pp 238
Thai LT, Maier WJ (1992) Solubilization abd biodegradation of octadecane in the presence of two commercial surfactants. Proceedings of the 47th Annual Purdue University Industrial Waste Conference, pp 167–175
U.S. Army Corps of Engineers (2003) Agriculturally based bioremediation of petroleum contaminated soils and shallow groundwater in Pacific island ecosystems
Van DK (1997) Fundamental of Petroleum, 4th Edition. Petroleum Extension Services, University of Texas at Austin, Austin, Texas
Van Beilen JB, Funhoff EG (2005) Expanding the alkane oxygenase toolbox: new enzymes and applications. Curr Opin Biotechnol 16:308–314
Van Beilen JB, Funhoff EG (2007) Alkane hydroxylases involved in microbial alkane degradation. Appl Microbiol Biotechnol 74:13–21
Van Beilen JB, Funhoff EG, Van Loon A, Just A, Kaysser L, Bouza M, Holtackers R, Röthlisberger M, Li Z, Witholt B (2006) Cytochrome P450 alkane hydroxylases of the CYP153 family are common in alkane-degrading eubacteria lacking integral membrane alkane hydroxylases. Appl Environ Microbiol 72:59–65
Van Dyke MI, Lee H, Trevors JT (1991) Applications of microbial surfactants. Biotech Adv 9:241–252
Velkov VV (2001) Stress-induced evolution and the biosafety of genetically modified microorganisms released into the environment. J Biosci 26:667–683
Venosa AD, Zhu X (2003) Biodegradation of crude oil contaminating marine shorelines and freshwater wetlands. Spill Sci Technol Bull 8:163–178
Verstraete W, Vanloocke R, DeBorger R, Verlinde A (1976) Modelling of the breakdown and the mobilization of hydrocarbons in unsaturated soil layers. In: Sharpley JM and Kaplan AM (eds) Proceedings of the 3rd International Biodegradation Symposium. Applied Science Publishers Ltd, London, p 99–112
Viswanathan S (1999) Bioremediation of water contaminated with petroleum hydrocarbons. Global environmental technology and management, Getm.Org
Volkering F, Breure AM, Van Andel JG (1993) Effect of microorganisms on the bioavailability and biodegradation of crystalline naphthalene. Appl Microbiol 40:535–540
Von Wedel RJ, Mosquera JF, Goldsmith CD, Hater GR, Wong A, Fox TA, Hunt WT, Paules MS, Quiros JM, Wiegand JW (1988) Bacterial biodegradation of petroleum hydrocarbons in groundwater: in situ augmented bioreclamation with enrichment isolates in California. Water Sci Technol 20:501–503
Walker JD, Colwell RR (1974) Microbial degradation of model petroleum at low temperatures. Microb Ecol 1:63–95
Walker JD, Colwell RR, Vaituzis Z, Meyer SA (1975) Petroleum-degrading achlorophyllous alga Prototheca zopfii . Nature 254:423–424
Walker JD, Colwell RR, Petrakis L (1976) Biodegradation rates of components of petroleum. Can J Microbiol 22:1209–1213
Ward DM, Brock TD (1976) Environmental factors influencing the rate of hydrocarbon oxidation in temperate lakes. Appl Environ Microbiol 31:764–772
Ward DM, Brock TD (1978a) Anaerobic metabolism of hexadecane in marine sediments. Geo Microbiol J 1:1–9
Ward DM, Brock TD (1978b) Hydrocarbon biodegradation in hypersaline environments. Appl Environ Microbiol 35:353–359
Weissenfel WD, Klewer HJ, Langhoff J (1992) Adsorption of Polycyclic Aromatic Hydrocarbons (PAH) by soil particles: influence on biodegradability and biotoxicity. Appl Microbiol Biotechnol 36:689–696
Wilkes H, Rabus R, Fischer T, Armstroff A, Behrends A, Widdel F (2002) Anaerobic degradation of n-hexane in a denitrifying bacterium: Further degradation of the initial intermediate (1- methylpentyl) succinate via C-skeleton rearrangement. Arch Microbiol 177:235–243
Youssef N, Simpson DR, Duncan KE, McInerney MJ, Folmsbee M, Fincher T, Knapp RM (2007) In situ biosurfactant production by Bacillus strains injected into a limestone petroleum reservoir. Appl Environ Microbiol 73:1239–1247
Yumoto I, Nakamura A, Iwata H, Kojima K, Kusmuto K, Nodasaka Y, Matsuyama H (2002) Dietzia psychralcaliphila sp. nov., a novel facultatively psychrophilic alkaliphile that grows on hydrocarbons. Int J Syst Evol Microbiol 52:85–90
Zengler K, Richnow HH, Rossello-Mora R, Michaelis W, Widdel F (1999) Methane formation from long-chain alkanes by anaerobic microorganisms. Nature 401:266–269
Zeyer J, Kuhn EP, Schwarzenbach RP (1986) Rapid microbial mineralization of toluene and 1,3-dimethylbenzene in the absence of molecular oxygen. Appl Environ Microbiol 52:944–947
Zimmer MT, Ohkuma A, Ohta M, Takagi Schunck W-H (1996) The CYP52 multigene family of Candida maltosa encodes functionally diverse n-alkane-inducible cytochromes p450. Biochem Biophys Res Commun 224:784–789
Download references
Author information
Authors and affiliations.
Department of Biotechnology and Allied Sciences, Jayoti Vidyapeeth Women’s University, Ajmer Road, Jaipur, Rajasthan, India
Subhash Chandra, Richa Sharma & Kriti Singh
Department of Biotechnology, Mahatma Gandhi Institute of Applied Sciences, Tonk Road, JECRC campus, Sitapura, Jaipur, Rajasthan, India
Anima Sharma
You can also search for this author in PubMed Google Scholar
Corresponding author
Correspondence to Subhash Chandra .
Rights and permissions
Reprints and permissions
About this article
Cite this article.
Chandra, S., Sharma, R., Singh, K. et al. Application of bioremediation technology in the environment contaminated with petroleum hydrocarbon. Ann Microbiol 63 , 417–431 (2013). https://doi.org/10.1007/s13213-012-0543-3
Download citation
Received : 02 April 2012
Accepted : 03 September 2012
Published : 21 September 2012
Issue Date : June 2013
DOI : https://doi.org/10.1007/s13213-012-0543-3
Share this article
Anyone you share the following link with will be able to read this content:
Sorry, a shareable link is not currently available for this article.
Provided by the Springer Nature SharedIt content-sharing initiative
- Environment
Annals of Microbiology
ISSN: 1869-2044
- Submission enquiries: [email protected]
- General enquiries: [email protected]
Bioremediation: a green approach for restoration of polluted ecosystems
- Published: 23 November 2018
- Volume 1 , pages 305–307, ( 2018 )
Cite this article
- Naveen Kumar Arora 1
13k Accesses
52 Citations
Explore all metrics
Avoid common mistakes on your manuscript.
Anthropogenic activities have resulted in deterioration of ecosystems throughout the globe making them unfit for survival of indigenous biological forms. This has also put a huge pressure on the fast receding natural resources. Environmental pollution has been a major concern over the past few decades influencing the quality of life. Rampant industrialization, improper agricultural methods, unchecked discharge of pollutants into land and water bodies has severely contaminated the ecosystems on earth. This has resulted in inadequate use of natural resources, increase in barren fields, loss of biodiversity, problem of potable water and huge economic losses which are very difficult to even estimate. Manmade chemicals are increasing by the day and many of them are recalcitrant and most being xenobiotic. As per estimates every year 10 million tons of toxic chemicals are released into the environment around the globe. Due to the addition of dangerous toxic chemicals such as polycyclic aromatic hydrocarbons (PAH), polychlorinated biphenyls (PCBs), soil and water systems have become contaminated. These pollutants are carcinogenic and are persistent, causing great harm to the ecosystems harming the health of the environment and causing damage to all life forms. Heavy metal pollution is another baleful influence of human activity on environment. Industrial processes including refineries, metal processing units, waste incineration, fossil fuel burning, nuclear power plants, lead based paints, plastics, electronic wastes, agrochemicals, pharmaceutical chemicals, vehicle exhausts, leather industries are the major sources of heavy metal contamination in the environment. Heavy metals degrade the quality of soil and water, rendering them unsustainable and of little or no use, also negatively impacting the biological health of ecosystems. The green revolution in 1950s or 60s although helped in increasing food productivity but caused acute toxification of the environment due to the injudicious use of chemical pesticides and fertilizers. Recalcitrant nature, particularly of the pesticides has resulted in their long term accumulation in soil, water and food chains. As per the report of Food and Agriculture Organization (FAO) production of pesticides has increased from 0.2 million tons in 1950 to 5 million tons in 2000 resulting in deterioration of fertile land, killing non-targeted microbes, birds, animals and proving fatal for human health. World Health Organization (WHO) estimates that pesticides cause quarter of a million premature deaths and over 3 million people are being hospitalized each year due to poisoning. Oil spills in oceans and on land have also played havoc with the respective ecosystems endangering the life and quality of natural systems to a great extent. In last 5 years 40,000 tonnes of oil was estimated to be spilled in the marine ecosystems through offshore mining, accidents, damaged tankers etc. The insoluble oil strata cuts off the oxygen supply and sunlight from entering the water body causing the death of flora and fauna, extinction of species and reduced microbial population. On land spilled oil forms a layer, impacting the oxygen levels and releasing toxic components, rendering the system non-fertile.
Apart from these, climate change caused by ever-increasing emission of green house gases (GHGs), has resulted in degradation of ecosystems due to rising temperature, droughts, floods etc. Incidences of heavy or very low rains have become common now throughout the globe causing flash floods or drought. Very recent floods in Kerala, India, one of the biggest in last 100 years in the region, have completely devastated the agricultural fields and yields due to accumulation of silt and washing off of the top soil. Similarly, wild fires in California have resulted in great loss to the forest, ecosystem of the region and also to human life, rendering the soil and the region greatly impacted. Incidences of heat waves in Europe and other parts of the world are now much more common. The impact of climate change and addition of chemicals has resulted in desertification and salinization of land resulting in lower productivity and lesser availability of biological resources. Approximately one billion hectares of land is suffering from the problem of salinity around the globe and this menace is increasing by the day. Water bodies including rivers, lakes or oceans have been impacted, causing great loss to their biodiversity or utility. With increasing human population we need more and more resources, productive fields, clean water bodies so as to get the maximum out of the natural resources. For this we have to clean-up the mess and remove toxic pollutants from the ecosystems, reclaim waste and marginal lands, saline soils, rejuvenate fresh water bodies and make oceans free of contaminants.
Over the years, various conventional methods like physical, chemical and thermal processes have been used to clean and remediate the ecosystems. But there have been several and some serious drawbacks associated with these processes, such as production of toxic intermediates, transport of contaminated soil/ water for treatment, high costs of treatment and inefficient revival of natural flora and fauna. However, use of biodegradation and bioremediation techniques involving biological systems such as microorganisms or their products and plants, are sustainable, cost effective options abating and rendering the pollutants harmless by natural biological activities. In bioremediation biological systems are applied for reclamation of the contaminated soil/ water by transformation of toxic pollutants into less hazardous or completely non-hazardous forms. The bioremediation technology includes extensive use of microorganisms or their enzymatic machineries, phytoremediation (plants) and rhizoremediation (plant and microbe interactions) techniques. Bioremediation is of two types depending on their site of application including cheaper and much effective in situ remediation where pollutants are treated on the site of contamination and the ex situ remediation where the contaminated samples are brought to laboratory and industries and are treated; used more for treatment of highly polluted but for smaller area or for systems such as ground water, where it is not advisable to add microbes or their enzymes. Technical aspects of bioremediation involve various mechanisms such as biosequestration, biodegradation, phytohydraulics, biological extraction, and volatilization by which microbes or plants immobilize or transform the complex moieties of the pollutants remediating the land and water. These biological systems have successfully been applied in cleaning up of ground water, lagoons, sludge, water streams, agricultural lands, oil spills, petroleum and hydrocarbon contaminated sites. Bioremediation through microorganisms generally involves the application of aerobic bacteria reported to degrade pesticides and hydrocarbons, both alkanes and polyaromatic compounds and several other pollutants. Anaerobic bacteria are used for bioremediation of PCBs in river sediments and dechlorination of trichloroethylene (TCE), and chloroform; lignolytic fungi for degradation of toxic and persistent pollutants; methylotrophs for remediation of chlorinated aliphatics trichloroethylene and 1, 2-dichloroethane and so on. Large scale treatment of petroleum hydrocarbon contamination in oceans, waste water treatment in polar regions, removal of toxic pollutants from agro-industrial wastes, treatment of polluted shorelines have been reported to be successfully done through microbial based bioremediation. Heavy metal contamination from soil is being effectively removed by microbial or phytoremediation techniques. Rhizoremediation is a cheap and efficient technique useful for remediation of contaminated soils by the combined action of plant and their symbiotic rhizosphere microbes. Plant growth promoting microbes have been used for reclamation of saline or non-fertile marginal lands by enhancing crop productivity. After repeated use year after year, these microbes with symbiotic partners i.e. plants help to reclaim the barren and saline soils making them fertile. This results in bioremediation of such wastelands and also control of climate change due to increased carbon sequestration by the remediated ecosystems. Many success stories of highly and vastly contaminated sites are now known, including long shore lines, as in Alaska oil spill, or decontamination of polluted soils or agriculture fields. Genetic engineering has very important role to play in the area of bioremediation. Capable microorganisms are engineered to improve their cell membrane transportation or their enzymatic attributes supporting the enhanced and wide spectral degradation of pollutants. “Superbugs” are the most popular examples of genetically engineered biological tools, significant in remediating oil spills and other toxic pollutants. Microbes are being engineered to have genes for degradation of multiple components of the complex crude oil and there are success stories in this area. Pyrosequencing, a next-generation molecular approach, is contributing to study pollutant-microbe interactions. With the help of pyrosequencing, environmental responses of microbes to contaminants, microbiome resistance against pollutants, diversity of fungal degrading genes in soil, can be known and used much more efficiently in bioremediation of contaminated sites. Quorum sensing properties of microbes have also been implemented for bioremediation as stimulation of signaling molecules like acylated homoserine lactone (AHL) also regulate gene expression for exopolysaccharides (EPS) and biofilm formation helping in degradation. Microbial fuel cells, microbial electrolysis cells and microbial desalination cells, bioelectrical wells and biofiltration are other advanced biotechnological approaches used for removal of various toxicants from the environment. Exploitation of effective genes from non-culturable microbes in a contaminated ecosystem can be used for degradation and remediation of complex or multiple polluted sites. Metagenomics, transcriptomics, metabolomics and fluxomics can play very important roles in future to identify the important genes involved in biodegradation, their expression, release of important enzymes and rates of metabolic reaction even in highly contaminated ecosystems. This can go a long way in improving the quality of stressed ecosystems, because most of them have huge populations of non-culturable microbes with great genetic pool and abilities to survive and flourish in such habitats. Need is to exploit them by the process of bioaugmentation of nutrients or through co-metabolism. Hence, although we have already achieved a bit in the area of bioremediation but a lot is yet to be explored and future of this green technology in cleaning up the environment is very bright.
Bioremediation has a great potential with notable achievements already reported from around the globe. But still this excellent and eco-friendly low input biotechnology has been underutilized. The global market scenario of bioremediation technology and services is showing an elevation with compound annual growth rate of 8.3% from 2017 to 2025. This can be much higher if exploited and developed properly. Bioremediation and biodegradation are the key focus areas of the journal “Environmental Sustainability”. We need to clean the mess created by anthropogenic activities through green technologies so as to provide a healthy and sustainable planet to the future generations.
Author information
Authors and affiliations.
Department of Environmental Science, BBA University, Lucknow, UP, India
Naveen Kumar Arora
You can also search for this author in PubMed Google Scholar
Corresponding author
Correspondence to Naveen Kumar Arora .
Rights and permissions
Reprints and permissions
About this article
Arora, N.K. Bioremediation: a green approach for restoration of polluted ecosystems. Environmental Sustainability 1 , 305–307 (2018). https://doi.org/10.1007/s42398-018-00036-y
Download citation
Published : 23 November 2018
Issue Date : 01 December 2018
DOI : https://doi.org/10.1007/s42398-018-00036-y
Share this article
Anyone you share the following link with will be able to read this content:
Sorry, a shareable link is not currently available for this article.
Provided by the Springer Nature SharedIt content-sharing initiative
- Find a journal
- Publish with us
- Track your research
An official website of the United States government
The .gov means it’s official. Federal government websites often end in .gov or .mil. Before sharing sensitive information, make sure you’re on a federal government site.
The site is secure. The https:// ensures that you are connecting to the official website and that any information you provide is encrypted and transmitted securely.
- Publications
- Account settings
- Advanced Search
- Journal List
- Appl Bionics Biomech
- v.2021; 2021

This article has been retracted.
Advances in heavy metal bioremediation: an overview.
1 Chemistry Department, Faculty of Applied Science, Umm Al-Qura University, Makkah, Saudi Arabia
Omar B. Ahmed
2 Department of Environmental and Health Research, The Custodian of the Two Holy Mosques Institute of Hajj and Umrah Research, Umm Al-Qura University, Makkah, Saudi Arabia
The pollution of toxic heavy metals is considered one of the most important environmental issues which has accelerated dramatically due to changing industrial activities. This review focuses on the most common methods, strategies, and biological approaches of heavy metal bioremediation. Also, it provides a general overview of the role of microorganisms in the bioremediation of heavy metals in polluted environments. Advanced methods of heavy metal remediation include physicochemical and biological methods; the latter can be further classified into in situ and ex situ bioremediation. The in situ process includes bioventing, biosparging, biostimulation, bioaugmentation, and phytoremediation. Ex situ bioremediation includes land farming, composting, biopiles, and bioreactors. Bioremediation uses naturally occurring microorganisms such as Pseudomonas , Sphingomonas , Rhodococcus , Alcaligenes , and Mycobacterium . Generally, bioremediation is of very less effort, less labor intensive, cheap, ecofriendly, sustainable, and relatively easy to implement. Most of the disadvantages of bioremediation relate to the slowness and time-consumption; furthermore, the products of biodegradation sometimes become more toxic than the original compound. The performance evaluation of bioremediation might be difficult as it has no acceptable endpoint. There is a need for further studies to develop bioremediation technologies in order to find more biological solutions for bioremediation of heavy metal contamination from different environmental systems.
1. Introduction
Heavy metals refer to metals with relatively high densities (more than 5gm/cm 3 ), atomic weights (greater than 50), and atomic numbers. They are often present in the earth as a normal component due to erosion process to rocks, naturally occurring decay of plant and animal waste matter, precipitation or atmospheric accumulation of airborne particles from volcanic eruption, and forest fire smoke. Moreover, wind erosion and oceanic spray also contribute to the exposure of heavy metals in the environment [ 1 ]. The pollution of toxic heavy metals is considered one of the most important environmental issues that has been accelerated dramatically due to changing industrial activities. Pollutants can be introduced and built up in the environment due to various human activities such as domestic waste, vehicles emission, industrial processes (e.g., electroplating, dyeing, and mining), the random disposal of electronic waste, agricultural fields, sewage sludge, and waste treatment plants [ 2 , 3 ]. In addition, heavy metals may be present at high levels in aquatic and soil ecosystems as compared with the atmosphere (e.g., vapors or particulate) [ 4 , 5 ]. The source of heavy metals may be either natural or due to human activities, which eventually leads to their presence in soil, water, and air [ 5 – 8 ] ( Figure 1 ). Contaminated soils and ground waters put human health at risk through the consumption of food grown on polluted areas, dermal contact, and the inhalation of dust [ 9 – 11 ]. Pollution by heavy metals is a serious threat to environmental living organisms depending on the concentration of heavy metal and the overdose absorbance rate. The negative impact of heavy metals to human and animal health is related to their long-term presence in the environment, and many heavy metals have high level of toxicity even at low concentrations (e.g., mercury, arsenic, lead, fluorine, and cadmium) [ 12 – 14 ]. Lead is considered one of the most persistent heavy metals because it can persist in soil up to 5,000 years with a high average biological half-life [ 15 ]. The main problem with heavy metals is the difficulty of biodegradation when bioaccumulation occurs in tissues. They are also capable of biomagnification along with the nutritional levels [ 16 ]. The discharge of heavy metals in the environment occurs through geological process (e.g., weathering and volcanic eruptions). Arsenic, lead, cadmium, nickel, chromium, cobalt, zinc, and selenium are highly toxic even in minor concentrations [ 17 , 18 ]. Remediation can be defined as the removal of pollution or contaminants from the environment (e.g., sediment, groundwater, and surface water and soil) to protect human health and restore the environment. This review discusses advances in heavy metal bioremediation from the existing literature by collecting databases and information from more than 60 publications that address the last issues related to bioremediation in the context of polluted environments. It also focuses on the most common methods, strategies, and biological approaches of heavy metal bioremediation and provides a general overview of the role of microorganisms in the bioremediation of heavy metals in polluted environments.

Sources and cycles of heavy metals in the environment [ 5 ].
2. Methods of Heavy Metal Remediation
In recent years, variable technologies and methods have been used in heavy metal remediation in contaminated environments such as soil and water. Such methods include physicochemical and biological methods; the latter is further classified into in situ and ex situ bioremediation.
2.1. Physicochemical Methods
Physicochemical methods include processes that work to remove heavy metals from any contaminated environment. They can be applied on the form of particulate of metals or metal-containing particles. This remediation can be conducted through physical and chemical processes such as ion exchange, precipitation, reverse osmosis, evaporative recovery, solvent extraction, filtration, chemical oxidation, chemical leaching, electrokinetics, land filling, electrochemical treatment, electrodialysis, ultrafiltration, solvent extraction, chemical precipitation, chemical reduction, and isolation (mechanical) separation of metals [ 1 , 9 , 19 , 20 ]. However, these methods may show incomplete metal removal; in addition, they require high solvent and production of poisonous waste products. They also have an inherent negative environmental impact and are usually soil disturbing, besides that fact that they are labor intensive and expensive [ 10 ]. So, these methods are limited by their high costs, high energy requirements, low efficiency, unpredictable metal ion removal, and generation of toxic sludge [ 21 – 27 ].
2.2. Biological Methods
Biological remediation or biodegradation constitutes many types of methods involved in the removal or degradation of heavy metals through biological activity. These biological treatments may either include aerobic (presence of oxygen) or anaerobic (absence of oxygen) processes and can be used for heavy metal removal. Biodegradation is a process wherein the polluted environment is biologically degraded under certain conditions to levels below the concentration limits established by regulatory authorities [ 28 – 32 ]. Bioremediation can further be classified into in situ or ex situ categories based on the strategies involved in Table 1 .
Types of bioremediation.
2.2.1. In Situ Bioremediation
In situ bioremediation methods treat the contamination at the site without removing soil. The use of these specific methods depends on many factors: the area contaminated, properties of the compounds involved, concentration of the contaminants, and time required to complete the bioremediation. This process is usually recommended because it requires moving fewer materials and is less expensive. In situ bioremediation is sometimes classified into intrinsic bioremediation and engineered in situ bioremediation. It includes so many types mainly bioventing, biosparging, biostimulation, bioaugmentation, and phytoremediation [ 33 – 36 ].
(1) Bioventing . Bioventing technique is the most commonly used in situ mechanism in which air and nutrients are supplied to polluted soil to stimulate the microorganisms (bacteria). Bioventing requires limited air flow and low oxygen rates to release pollutants to the atmosphere through biodegradation. It can simulate in situ biodegradation of simple hydrocarbons in the soil, and hence, the contamination occurs deep under the surface [ 37 ]. Bioventing is limited by the inability to deliver oxygen to the polluted soil and the insufficient aeration of shallow contamination [ 38 ].
(2) Biosparging . Biosparging is defined as the injection of law pressure air below the water table to raise the level of groundwater oxygen and enhance the rate of bacterial bioremediation of contaminants [ 33 ]. Both bioventing and biosparging techniques have been utilized simultaneously to guarantee the efficient removal of the soil contaminants despite any unfavorable conditions. Biosparging can also combine soil and groundwater to reduce the concentration of dissolved oil compounds in groundwater, mixed with soil under level under water table and within the capillary fringe. It is an ease and low-cost procedure with considerable flexibility.
(3) Biostimulation . Biostimulation involves enhancing bacterial growth to initiate the bioremediation process. First, the polluted soil is mixed with enriched nutrients and vital substances to stimulate microbial activity for the fast degrading of contaminants or toxic compounds into the carbon source or nitrogen and phosphorous source [ 39 ]. Microorganisms such as bacteria and fungi represent nature's original recyclers. The capability of microorganism to transform chemical pollution into sources of energy and useful materials suggests important biological processes which are lower in cost and friend to environment.
(4) Bioaugmentation . In bioaugmentation, there are certain sites where microorganisms are required to extract the contaminants. They are also able to outcompete indigenous microorganisms, which means that they can clean up the site rapidly. The removal of toxic chemicals through bioaugmentation has been reported in environments such as soil and water. However, a number of limitations have also been documented. For instance, it has been observed that there is a decrease in the number of exogenous microorganisms after their addition to a polluted site due to abiotic and biotic stresses. They occur due to insufficient growth nutrients such as substrates, temperature changes, and pH, in addition to the competition between introduced and indigenous microorganisms [ 40 , 41 ].
(5) Phytoremediation . Phytoremediation represents an emerging technology that uses plants to remove pollution from soil and water. It has a potential use in the biodegradation of organic contaminants and may be a promising choice in the future. This technology is suitable for sites with shallow contaminants. Nevertheless, many studies have highlighted several limitations of this technology, such as contamination concentration, toxicity, bioavailability, the type of plant, and stress tolerance [ 42 ].
2.2.2. Ex Situ Bioremediation
Ex situ bioremediation means excavating and treating soil prior to returning the soil to its original state. If the contaminated material is excavated, it can be treated on or off site, which is often a more rapid method of decontaminating the area. Ex situ bioremediation is categorized into solid phase and slurry phase systems. The most important techniques include land farming, composting, biopiles, and bioreactors [ 33 ].
(1) Land Farming . Land farming is a simple process which implies that excavation of contaminated soil over a prepared site with periodic tilling until pollutants is degraded through microorganisms where the practice is limited to the treatment of small part of soil [ 43 ]. The technique is easy and much effective specially when used for petroleum-contaminated soil. However, the technique is limited to the treatment of a small space of upper soil (10–35 cm).
(2) Composting . A composting method involves combining polluted soil with nonhazardous organic agricultural wastes to support the growth of high microbial number with increased temperatures (40-65°C). The method is applied to a mixture of excavated soils and biosolids (wood chips, animal, and vegetative wastes) contaminated with organic materials (petroleum hydrocarbons and pesticides) [ 33 , 43 ].
(3) Biopiles . Biopiles are a mix or hybrid between land farming and composting. Biopiles produce enriched environments for different microorganisms (both aerobic and anaerobic). The aqueous reactors represent an ex situ treatment of a contaminated environment, where reactors are pumped up from a certain site. It involves bioremediating a contaminated environment by using a special engineered technology [ 33 ]. Engineered cells are made for the treatment of surface pollutants to regulate physical losses of the contaminants through leaching, which is then followed by volatilization [ 43 ]. Biopiling is considered a feasible, cost-effective technique for contaminated soils.
(4) Bioreactor . A bioreactor is a vessel used after a certain optimization of an external environment in which a biochemical reaction occurs. The system may include enzymes, tissues, microorganisms, and animal and plant cells to achieve a high yield of bioremediation. Overall, biodegradation is higher in bioreactor systems as compared with other systems because the target environment is easy to manage, control, and predict. Despite the advantages of reactor systems, it is found that the contaminated environment (e.g., soil) requires excavation of the contaminant from the soil through physical extraction before being processed by a bioreactor [ 33 ].
3. Role of Microorganisms
Various species of microorganisms can be used for bioremediation, as they are nature's original recyclers. They are also capable of transforming chemicals into sources of energy and raw materials for their own growth to produce a low-cost and environment-friendly biological process. Due to their high industrial use, heavy metals have become a worldwide real environmental problem. Toxic heavy metals are accumulated through the food chain due the industrial activities and fuel consumption leading to both environmental and health problems. These heavy metals exert toxic effects on living cells (mercury, silver, lead, cadmium, and arsenic). Many types of bacteria carry resistant genes to many types of cations and oxyanions of heavy metals in their DNA. To survive, bacteria undergo many different mechanisms to face the uptake of heavy metal ions. These mechanisms include biosorption, entrapment, efflux, reduction, precipitation, and complexation. Microorganisms therefore can be a promising, unlimited resource for new environmental biotechnologies. Bioremediation uses naturally occurring microorganisms to degrade or detoxify hazardous substances to human health and/or the environment. The microorganisms can either be used indigenously or be isolated from other resources at the polluted site [ 44 , 45 ]. Microorganisms which are involved in biodegradation are shown in Table 2 , including the following examples: Acinetobacter , Actinobactera , Alcaligenes , Arthrobacter , Bacillins , Beijerinckia , Flavobacterium , Methylosinus , Mycobacterium , Mycococcus , Nitrosomonas , Nocardia , Xanthobacter , Penicillium , Phanerochaete , Pseudomonas , Rhizoctonia , Trametes , and Serratia [ 46 – 66 ]. Most bioremediation processes are completed under aerobic conditions, but running a system under anaerobic conditions may permit microbial organisms to degrade otherwise recalcitrant molecules [ 44 ]. Aerobic organisms depend on oxygen during their growth activity. These are continuous processes which are known as cellular respiration, which use oxygen to oxidize substrates like fatty acid from oil in order to obtain energy. Examples of degradative aerobic bacteria are Pseudomonas , Sphingomonas , Rhodococcus , Alcaligenes , and Mycobacterium . Microorganisms can also be used to degrade toxic chemicals such as pesticides besides hydrocarbons materials [ 33 ]. Many bacteria use the contaminant as a metabolic source (carbon and energy). An anaerobic bacterium is an organism that does not need oxygen as a based metabolism, and it differs from aerobic bacteria. Anaerobic bacteria have also been used for the bioremediation of biphenyls, dechlorination, and chloroform [ 33 ]. Furthermore, fungi may be able to degrade a high range of persistent or toxic environmental pollutants [ 45 ]. There are many types of substrates available, such as corn cobs, straw, and dust. The aerobic bacteria that grow by utilizing methane for carbon and energy. This aerobic degradation being initiated with enzyme methane monooxygenase will be active against a wide range of chemicals [ 33 ].
Heavy metal distribution in environment and microorganisms involved in biodegradation.
As: arsenic; Cd: cadmium; Cr: chromium; Pb: lead; Hg: mercury; Cu: copper; Zn: zinc; Ni: nickel; Co: cobalt.
4. Factors Affecting Bioremediation
The removal of heavy metals through microorganisms may have ecological and economic limits. Several factors should be considered for the selection of a proper bioremediation. There are certain variables that have a great impact on the extent of biodegradation. First, nutrients in the polluted environment such as nitrogen, phosphate, sulphur iron, and potassium can stimulate and support strong microbial growth, cellular metabolism, and microorganism growth [ 34 ]. These nutrients represent basic life requirements and help microorganisms produce necessary enzymes to break down contaminants. Second, the remediation cost may also play an important role in the continuity of bioremediation, meaning that the cost should be low for financial feasibility. Third, the nature of pollutants, meaning whether they are solid, semisolid, liquid, or volatile in nature, may affect the process, or the pollutants are toxic or nontoxic, organic, and inorganic pollutants, heavy metals, polycyclic aromatic hydrocarbons, pesticides, and chlorinated solvents. The nature of the polluted area is also highly important, as it can affect the quality of bioremediation. Fourth, pH, temperature, and other physicochemical factors are important for the bioremediation process. The selection of the optimum range of these parameters can also greatly influence the rate and extent of biodegradation, as it influences the microbial growth and hence the removal of the contaminants [ 33 ]. Fifth, moisture content (water) is also a primary factor for biological growth and efficient bioremediation. Sixth, microbial diversity that can biodegrade any contaminant such as Pseudomonas, Aeromonas, Flavobacteria, Aeromonas, Chlorobacteria, Corynebacteria, Acinetobacter, Mycobacteria, Streptomyces, Bacilli, Macrobenthos, and other aquatic plants such as E. crassipes and L. hoffmeisteri can also degrade turbidity and chemical domestic wastewater [ 35 ]. Seventh, oxygen is mainly used for the initial breakdown of the hydrocarbon in the contaminated sites and also can be used for both aerobic and anaerobic bioremediation [ 36 ].
5. Advantages and Limitations
Bioremediation is a simple process used by many scientists in the waste treatment process for contaminated environments such as soil. The microbes that degrade the contaminant increase in numbers and release harmless products. The residues for the treatment are usually harmless products such as carbon dioxide, water, and cell biomass. Bioremediation is of very less effort, less labor intensive, and cheap compared to other methods that are used for the removal of hazardous waste. Bioremediation is also ecofriendly, sustainable, and relatively easy to implement. It is also useful for the complete destruction of a wide variety of contaminants [ 64 ]. Many hazardous compounds can be transformed into harmless products. Moreover, bioremediation can be implemented on the site of contamination itself without causing a major disruption of normal activities. There is no need to transport large numbers of waste off-site, there is no potential human health risk, and the environment will remain uncontaminated. Most of the disadvantages of bioremediation relate to it needing a longer time to be completed as compared with other options such as excavation and removing pollutants from the site. Also, there is a difficulty of bioremediation in treating inorganic contaminants and in confirming whether contaminants have been destroyed or not. Besides that, there is a slowness of highly chlorinated materials biodegradation and generation of more toxic or carcinogenic by-products [ 65 , 66 ]. Furthermore, the products of biodegradation sometimes become more toxic than the original compound. Its biological processes are also highly specific. Examples for effective site factors include the presence of microbial populations, growth conditions, and quantity of nutrients and pollutants [ 33 , 65 ].
6. Conclusion
Bioremediation technique is still a useful, natural, and environmentally friendly process in which the polluted environment is biologically biodegraded. Microorganisms play a significant role in the removal of heavy metals pollutants. The heavy metals (e.g., mercury, silver, lead, cadmium, and arsenic) exert toxic effects on living cells. Examples of degradative aerobic bacteria are Pseudomonas , Alcaligenes , Sphingomonas , Rhodococcus , and Mycobacterium . Anaerobic bacteria have also been used for the bioremediation of biphenyls, dechlorination, and chloroform. Furthermore, fungus microorganisms can effectively degrade many toxic environmental pollutants. Phytoremediation represents an emerging technology through which plants can be used to remove pollution from soil, water, and other environments. Bioremediation is of very less effort, less labor intensive, cheap, ecofriendly, sustainable, and relatively easy to implement. Most of the disadvantages of bioremediation relate to the slowness and time-consumption; furthermore, the products of biodegradation sometimes become more toxic than the original compound. Bioremediation may be limited by irregularity and uncertainty of completeness. Also, the performance evaluation of bioremediation might be difficult as there is no acceptable endpoint. There is a need for further studies to develop bioremediation technologies in order to find more biological solutions for bioremediation of heavy metal contamination from different environmental systems.
Acknowledgments
The authors would like to thank the Deanship of Scientific Research at Umm Al-Qura University for the continuous support. This work was supported financially by the Deanship of Scientific Research at Umm Al-Qura University (Grant Code: 18-SCI-1-02-0002).
Conflicts of Interest
The authors declare that they have no conflicts of interest.

VIDEO
COMMENTS
Bioremediation is a process that uses living organisms, mostly microorganisms and plants, to degrade and reduce or detoxify waste products and pollutants. ... Research Highlights 14 Dec 2022 ...
of the bioremediation research and knowledge, that is, heavy metal pollution, ... biotechnology and publication includes 70 full length research papers in international and national journals of .
Bioremediation research has largely focused on bacterial processes, which have numerous applications. ... Omics has gained prominence in the field of microbial remediation of the pulp and paper industry, textile industry, food industry, dairy industry, wood industry, fisheries, water and soil treatment industry, solid waste remediation, heavy ...
According to the United States National Research Council (US NRC), there are three criteria that must be met in intrinsic bioremediation and these include: demonstration of contaminants loss from contaminated sites, demonstration based on laboratory analyses that microorganisms isolated from contaminated sites have the innate potentials to ...
Authors' contributions cover various topics with a range of papers including original research and review articles spanning studies in remediation of different environments which outline new findings in the biotechnology field. This special issue contains five papers including one review article and four original research articles.
Bioremediation is the most effective management tool to manage the polluted environment and recover contaminated soil. Bioremediation, both in situ and ex-situ have also enjoyed strong scientific ...
Bioremediation Journal is a peer-reviewed quarterly that publishes current, original laboratory, bench, pilot-scale, and field research in biodegradation and bioremediation, the use of biological and supporting physical treatments, and biotechnolgies to treat contaminated soil, sediment, surface and groundwater. The journal rapidly disseminates new information on emerging and maturing ...
Though bioremediation is eco-friendly and economical for restoring the biological and physicochemical properties of the degraded soil (Arora 2018; Guang-Guo 2018), but still these in-situ techniques need to be improvised and more research and ethical issues need to be tackled for the use of improvised and efficient GEM's.
Bioremediation employs microbial metabolism under optimum environmental conditions and sufficient nutrients to degrade contaminants. This has proven to be effective and reliable due to its eco-friendly nature. Both in situ and ex situ techniques of bioremediation can be employed to reduce pollutant concentration.
Earlier published research papers have emphasized on individual genomic approaches in-depth for microbe mediated bioremediation. Hence, the present review aims to discuss a brief overview of the prevalent multi-omics techniques to investigate and understand the structural and functional aspects of the microbial community involved in bioremediation.
The algal bioremediation and biosynthesis of nanoparticles are two aspects of the same process that occur in parallel (Dahoumane et al., 2016). Transformation from bioremediation to biosynthesis can be observed using transmission electron microscopy (TEM), dynamic-light scattering (DLS), UV-vis spectroscopy, and scanning electron microscopy ...
This paper aims to describe the current bioremediation technologies, potential microorganisms involved, recent bioremediation treatment research to solve environmental damage and economic ...
Bioremediation Journal, Volume 27, Issue 4 (2023) See all volumes and issues. Volume 27, 2023 Vol 26, 2022 Vol 25, 2021 Vol 24, 2020 Vol 23, 2019 Vol 22, 2018 Vol 21, 2017 Vol 20, 2016 Vol 19, 2015 Vol 18, 2014 Vol 17, 2013 Vol 16, 2012 Vol 15, 2011 Vol 14, 2010 Vol 13, 2009 Vol 12, 2008 Vol 11, 2007 Vol 10, 2006 Vol 9, 2005 Vol 8, 2004 Vol 7 ...
Bioremediation is an emerging technology which can be simultaneously used with other physical and chemical treatment methods for complete management of diverse group of environmental pollutants. It seems as a sustainable approach for the environmental pollution management, and hence, there is a need for more research in this area. Efforts need ...
Nano-enhanced bioremediation is a new field of study, and more research is undeniably needed. The interdisciplinary combination offers new opportunities and novel functionalities such as. 1. Sustainable and environmentally friendly materials that can be used as alternatives to traditional post oil spill response.
The bioremediation studies after the Exxon Valdez oil spill demonstrated the promise of this technology for cleaning up a petroleum-contaminated site (NOAA Technical Memorandum 1997). Bioremediation was more extensive in 1990, with 378 of the 587 shoreline segments treated that year receiving bioremediation application (Chianelli et al. 1991 ...
Furthermore, it is challenging to extrapolate bench or pilot-scale operations and research to large-scale site operations and research in order to successfully construct the process. A few issues have also been highlighted about the toxicity of end products in the process of bioremediation when compared to the untreated parent compound ( Amin ...
bioremediation comprises basic research areas (cometabol ism, biotransformation kinetics, biotreatment, and biogeochemical modeling) and fie ld application research areas (biogeochemical ...
Bioremediation reduces pesticide contamination of agricultural soils by biodegradation processes via the metabolic activities of microorganisms. It is an efficient, cost-effective, and environment-friendly treatment. ... Unfortunately, this is mainly due to the poor cooperation among research laboratories, local authorities imposing a given ...
Bioremediation has a great potential with notable achievements already reported from around the globe. But still this excellent and eco-friendly low input biotechnology has been underutilized. The global market scenario of bioremediation technology and services is showing an elevation with compound annual growth rate of 8.3% from 2017 to 2025.
Bioremediation technologies, on the other hand, ... Thus, more research is needed on the identification/isolation of autochthonous bacterial strains with high petroleum biodegradation potential to increase the range of native bacterial consortia products for hydrocarbons bioremediation applications. Moreover, the procedures needed to fully use ...
In this study, we focused on soil contaminated by polycyclic aromatic hydrocarbons (PAHs) at typical coking-polluted sites in Beijing, conducted research on enhanced PAH bioremediation and methods to evaluate remediation effects based on toxicity testing, and examined changes in pollutant concentrations during ozone preoxidation coupled with biodegradation in test soil samples. The toxicity of ...
Abstract. This paper presents a brief outline of the development of bioremediation technologies (Microorganism remediation, phytoremediation, animal remediation and other methods applied in ...
Also, it provides a general overview of the role of microorganisms in the bioremediation of heavy metals in polluted environments. Advanced methods of heavy metal remediation include physicochemical and biological methods; the latter can be further classified into in situ and ex situ bioremediation. The in situ process includes bioventing ...